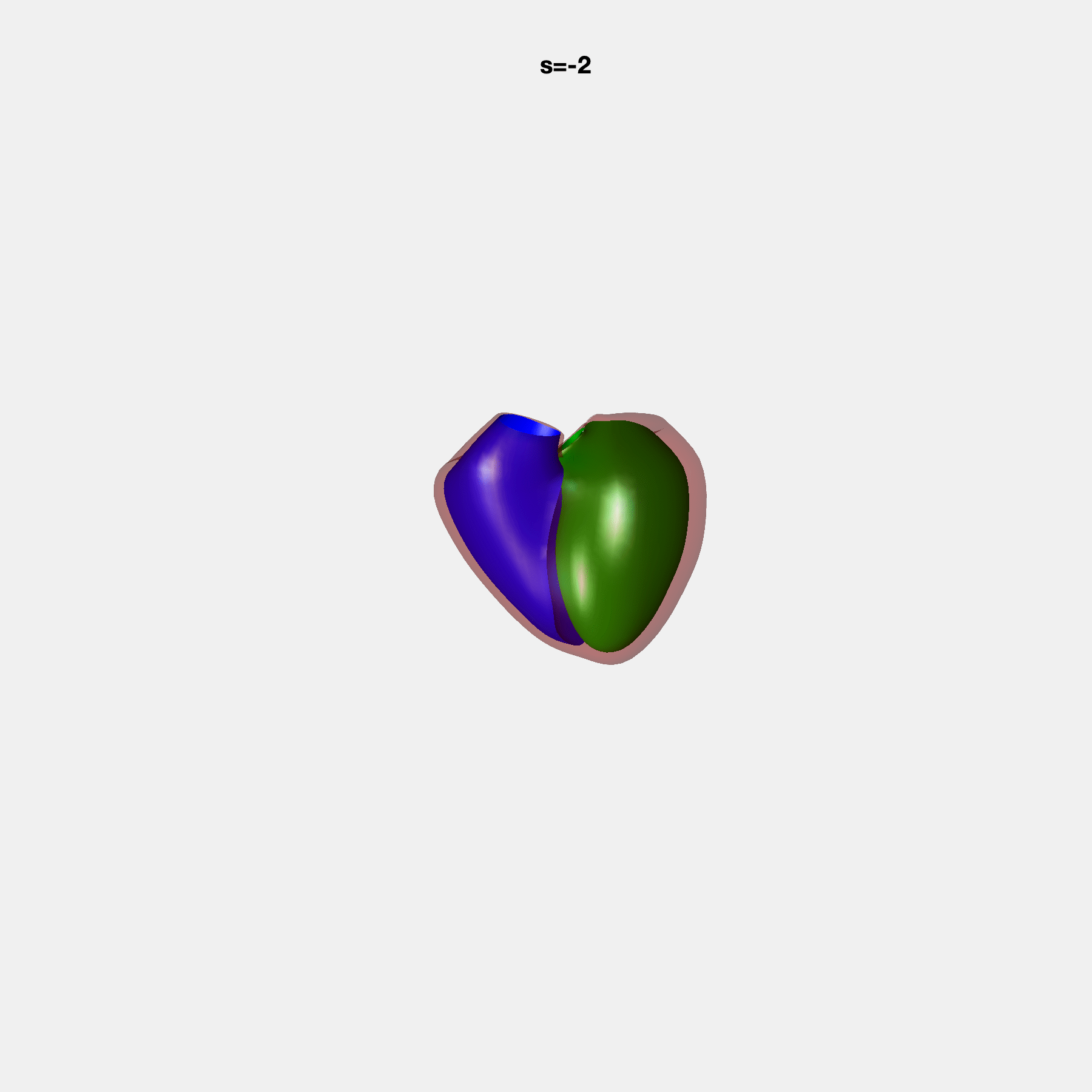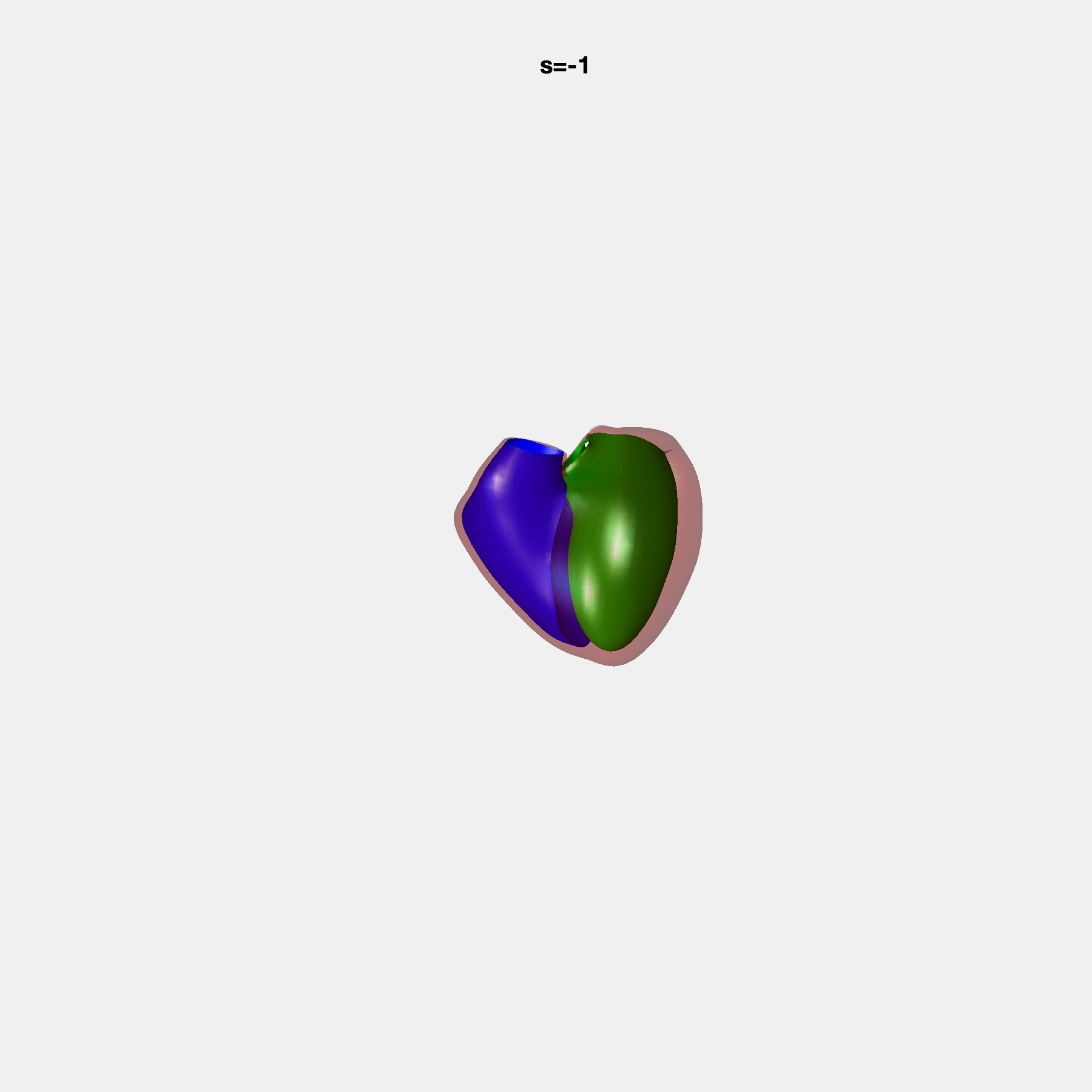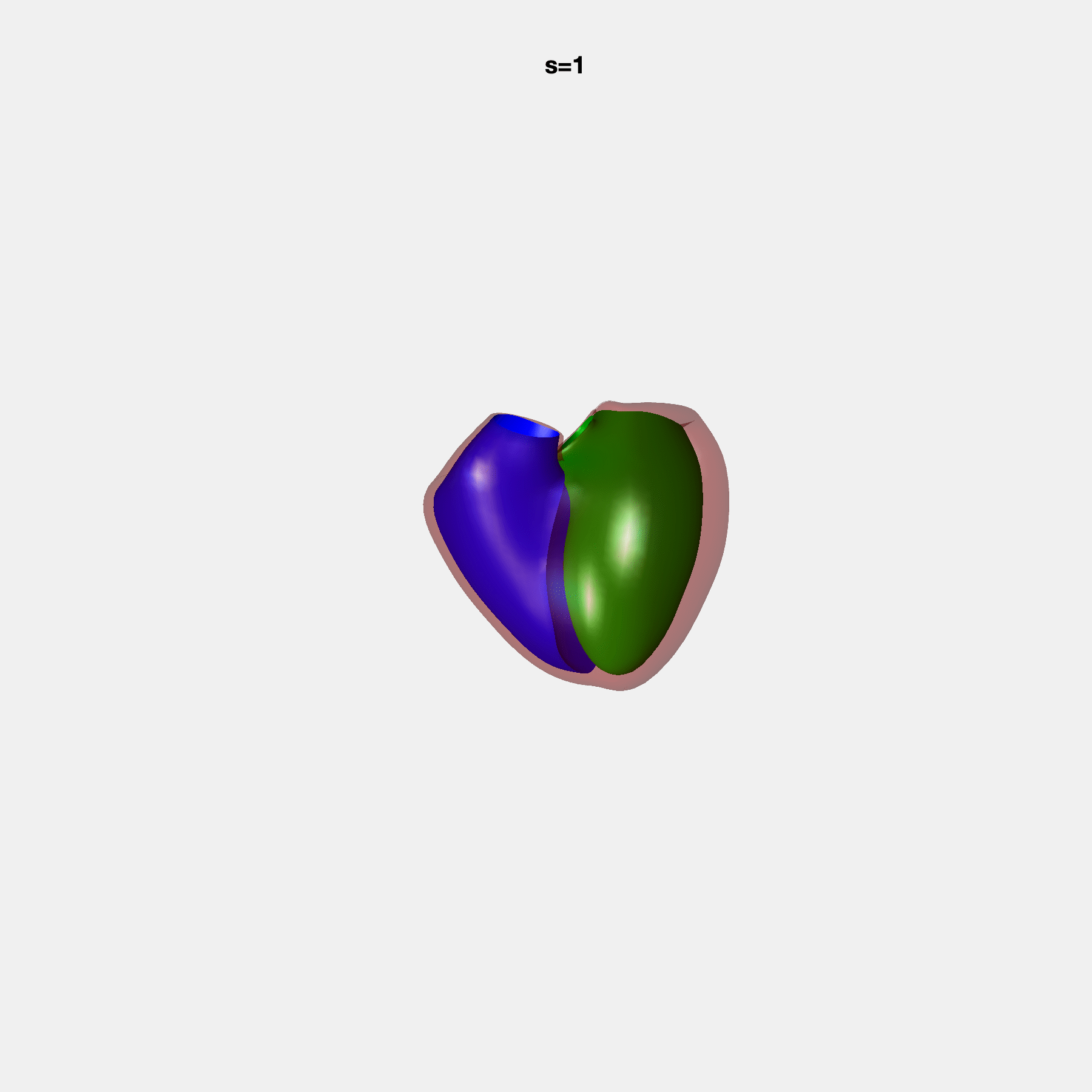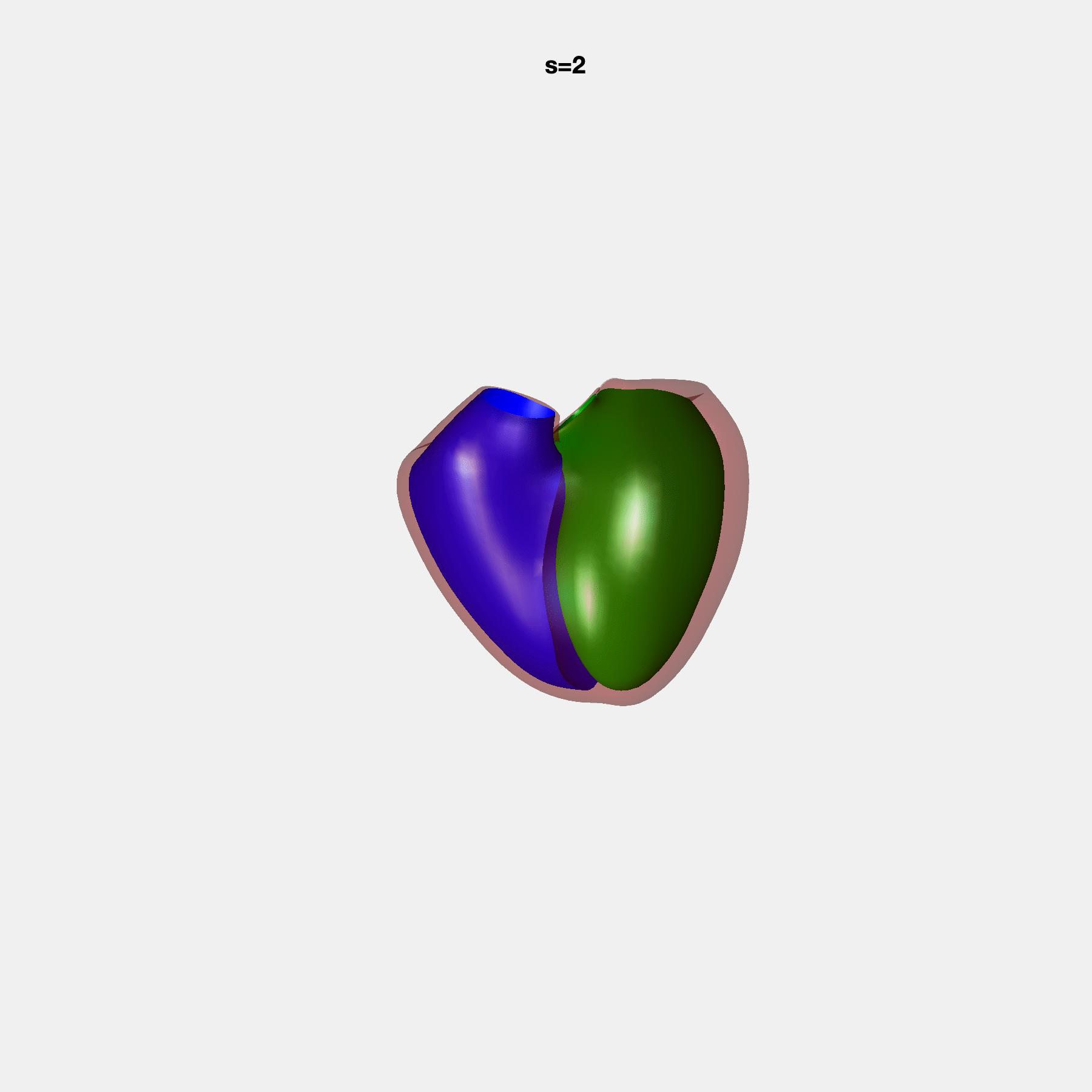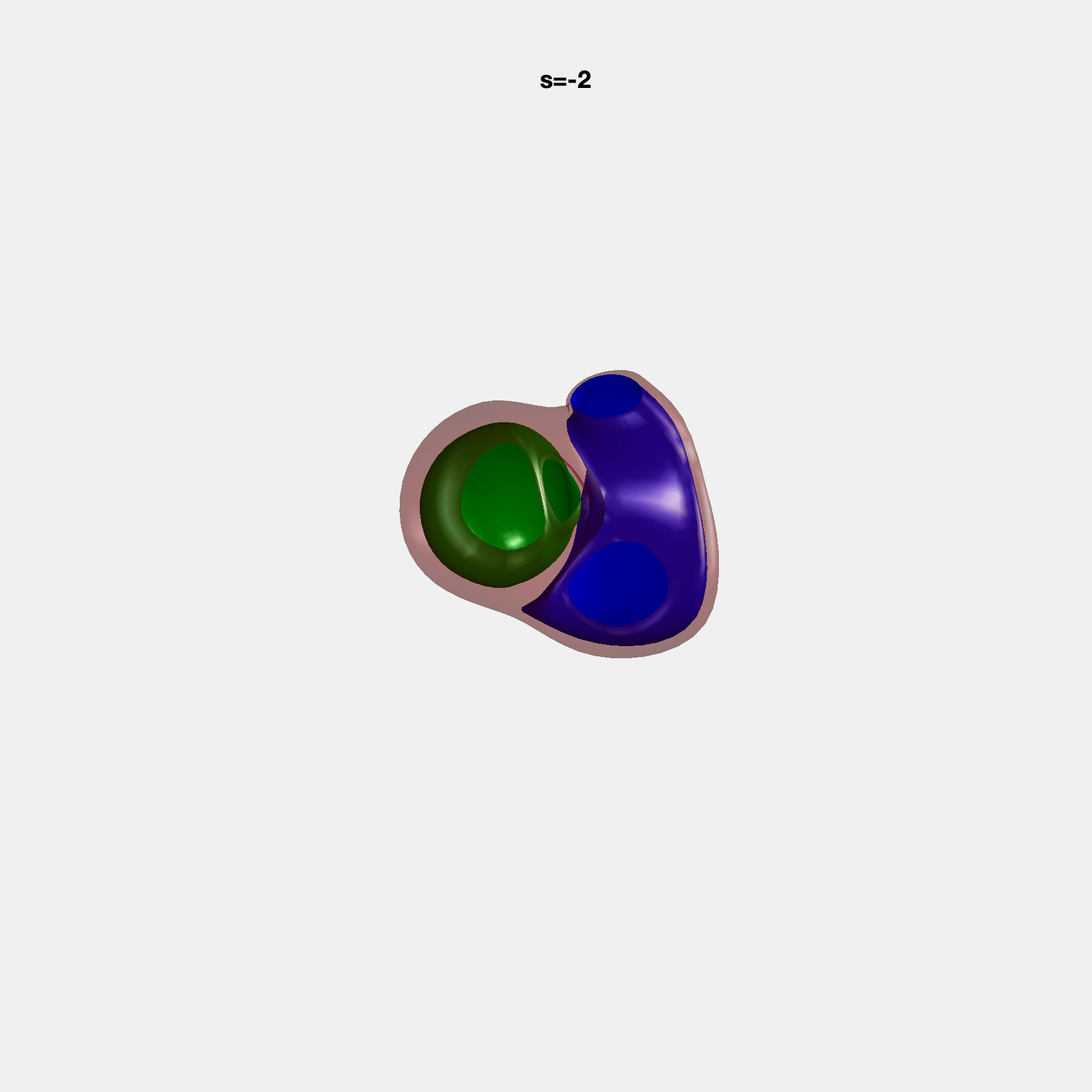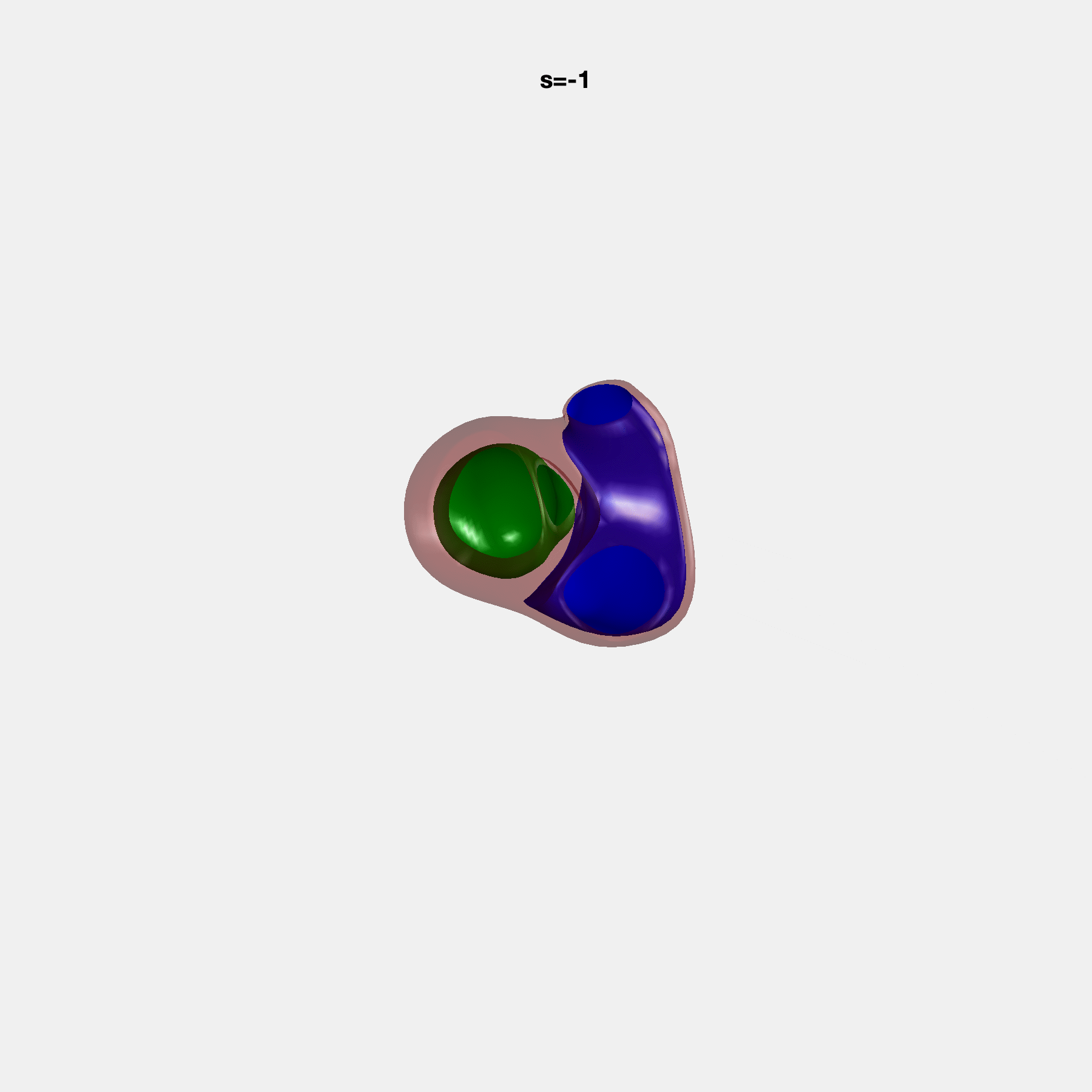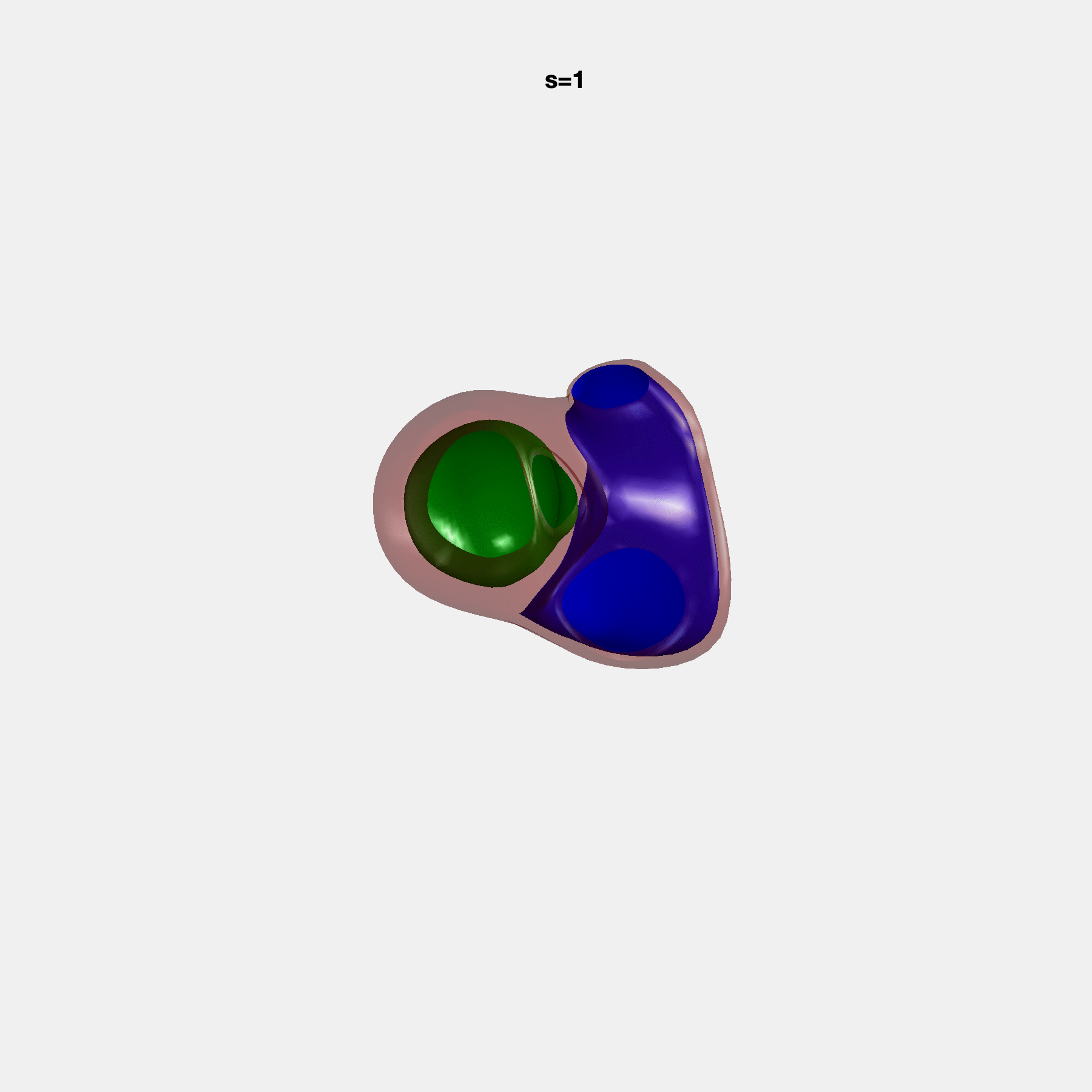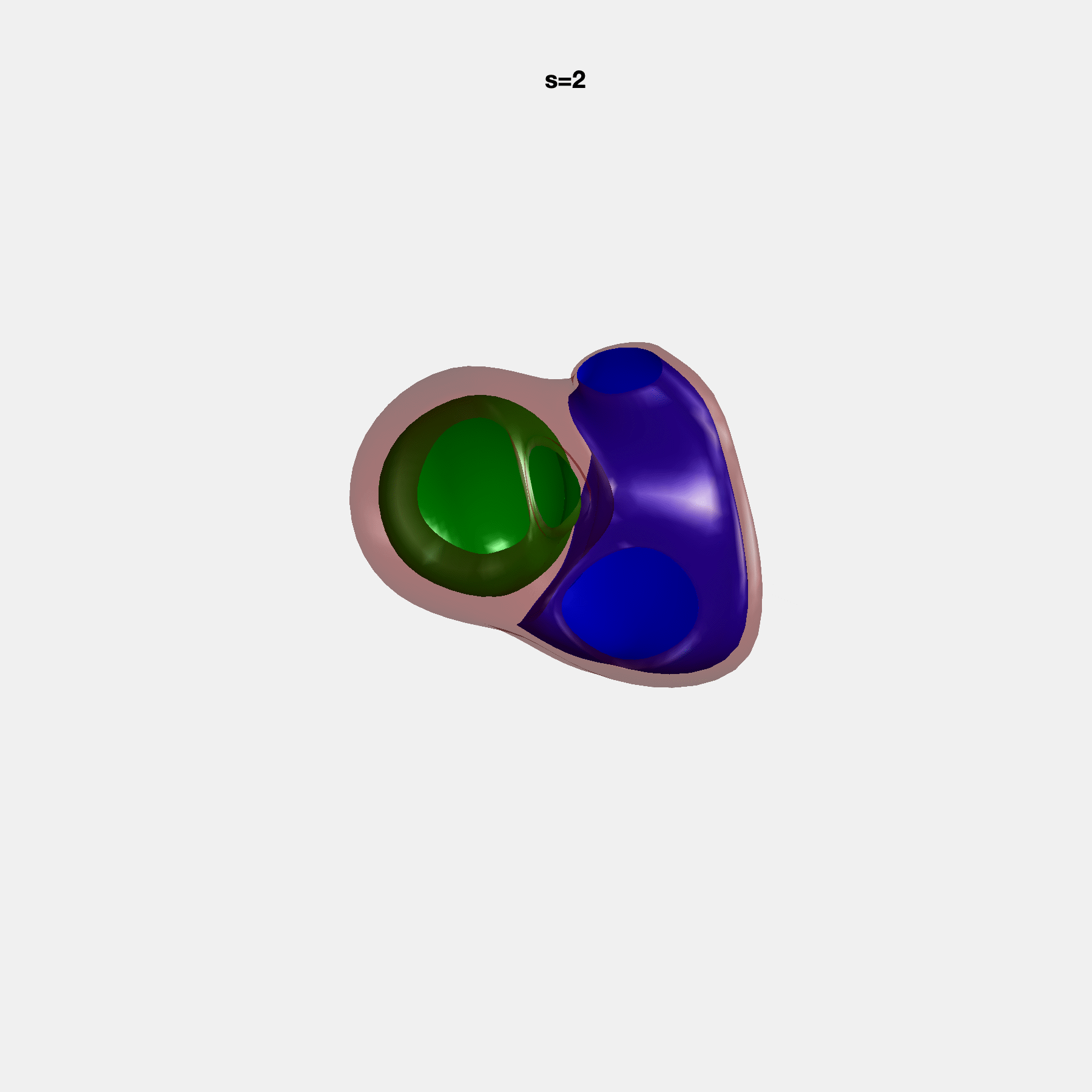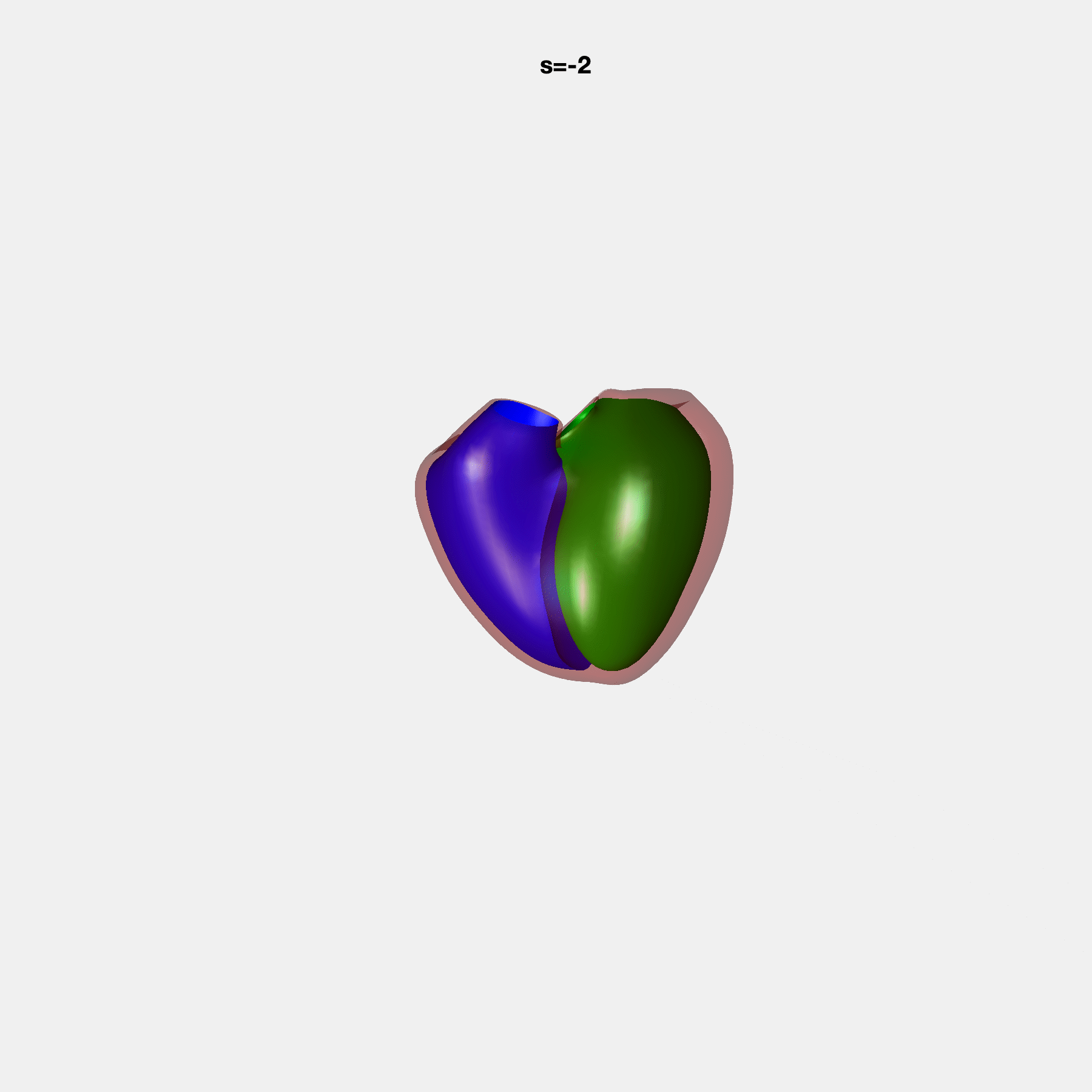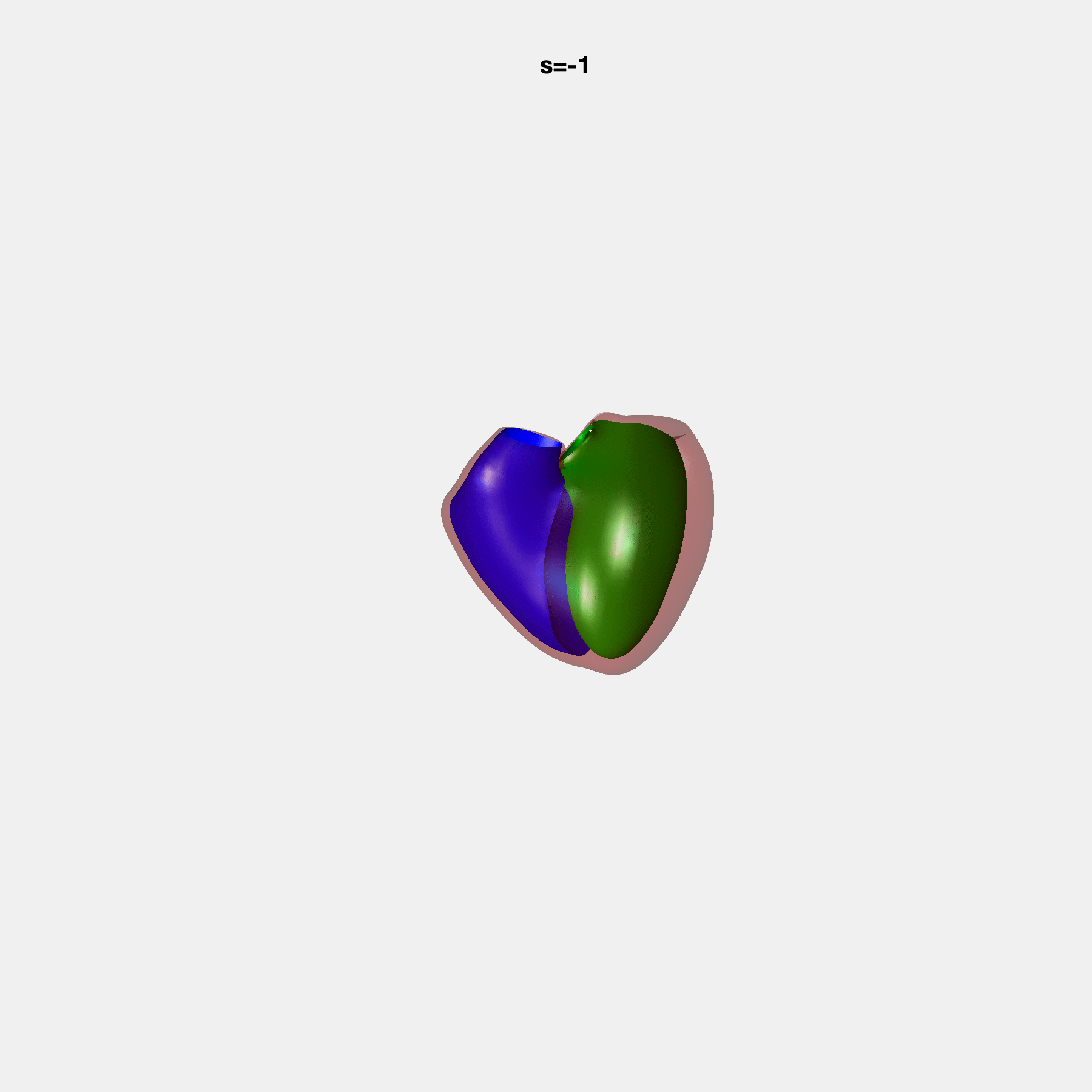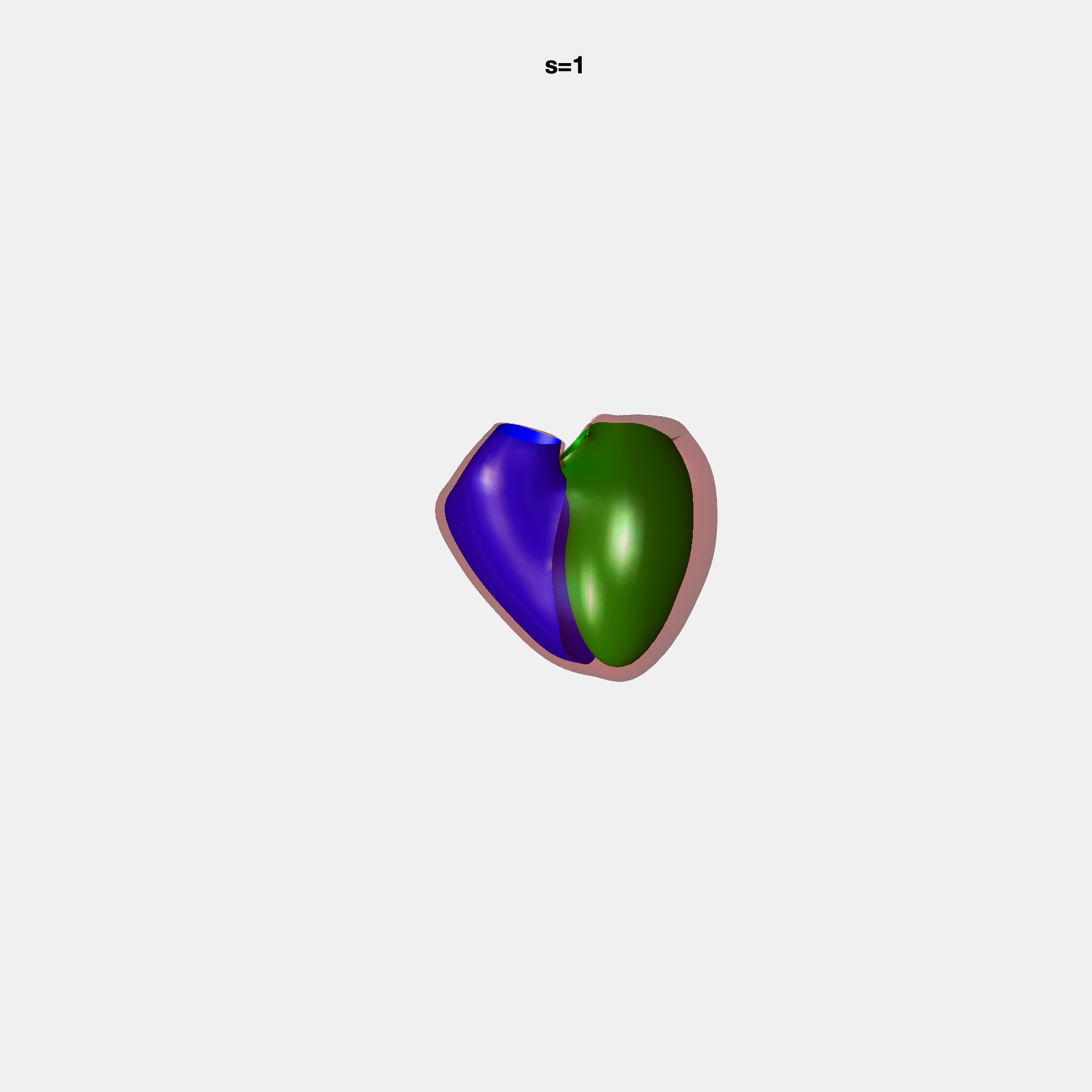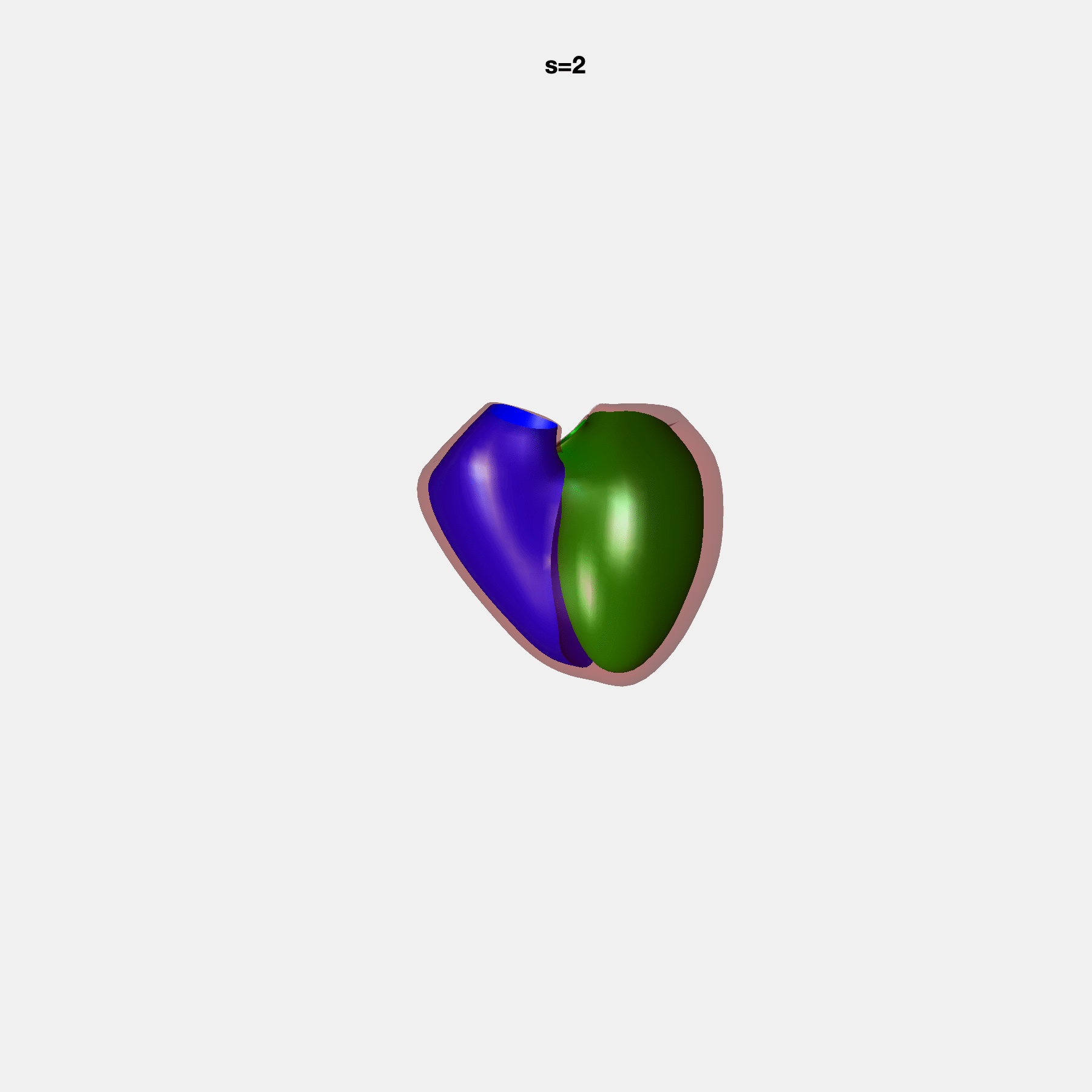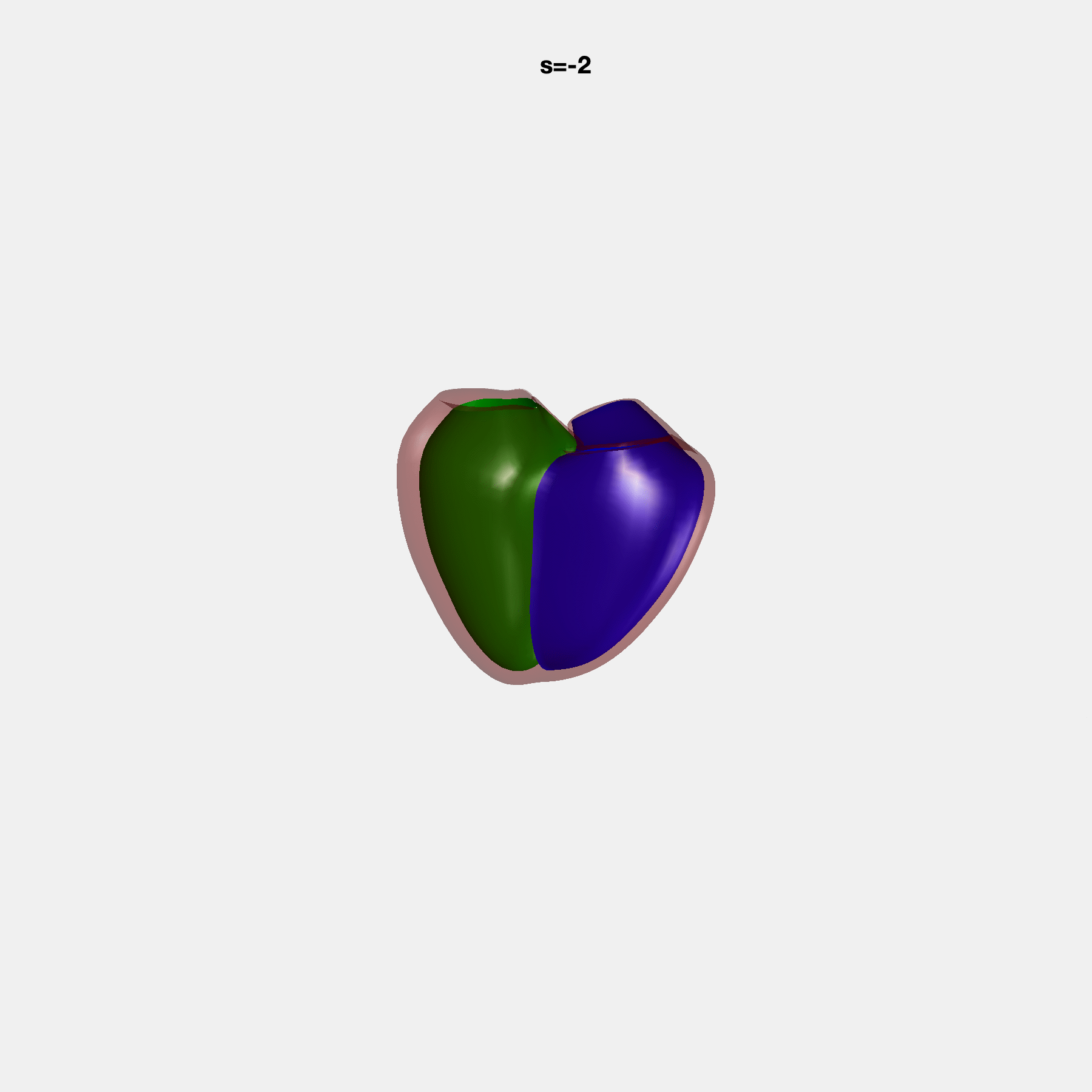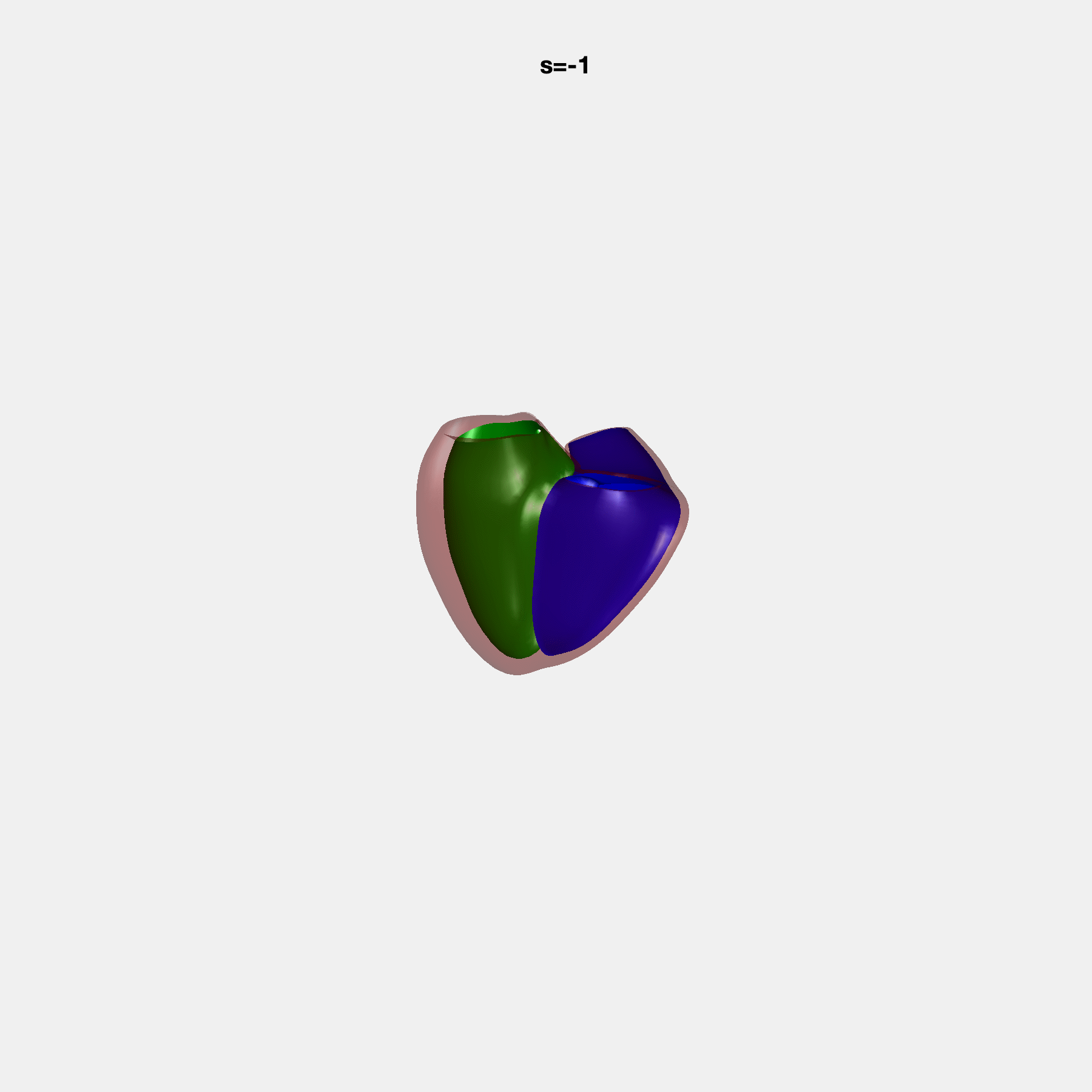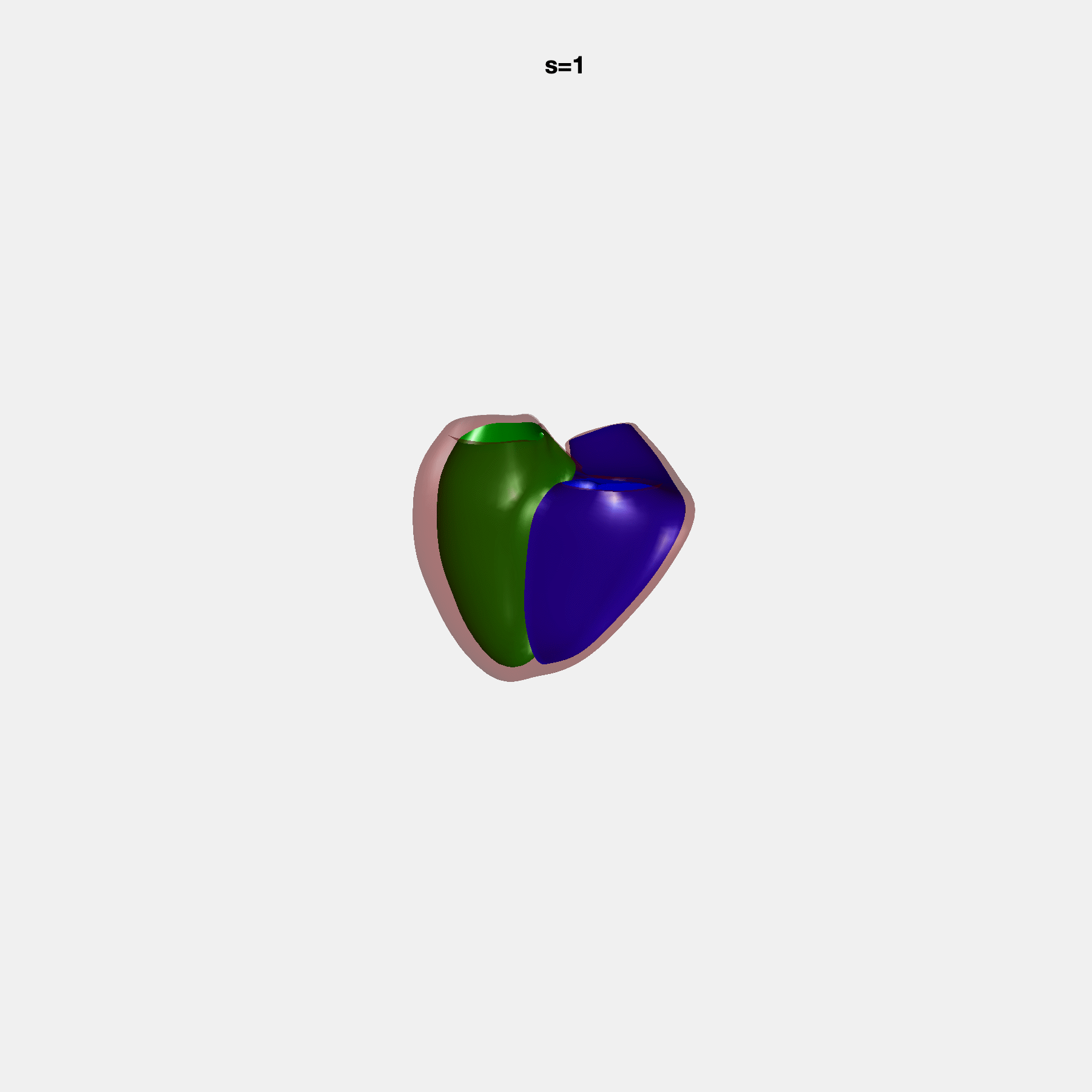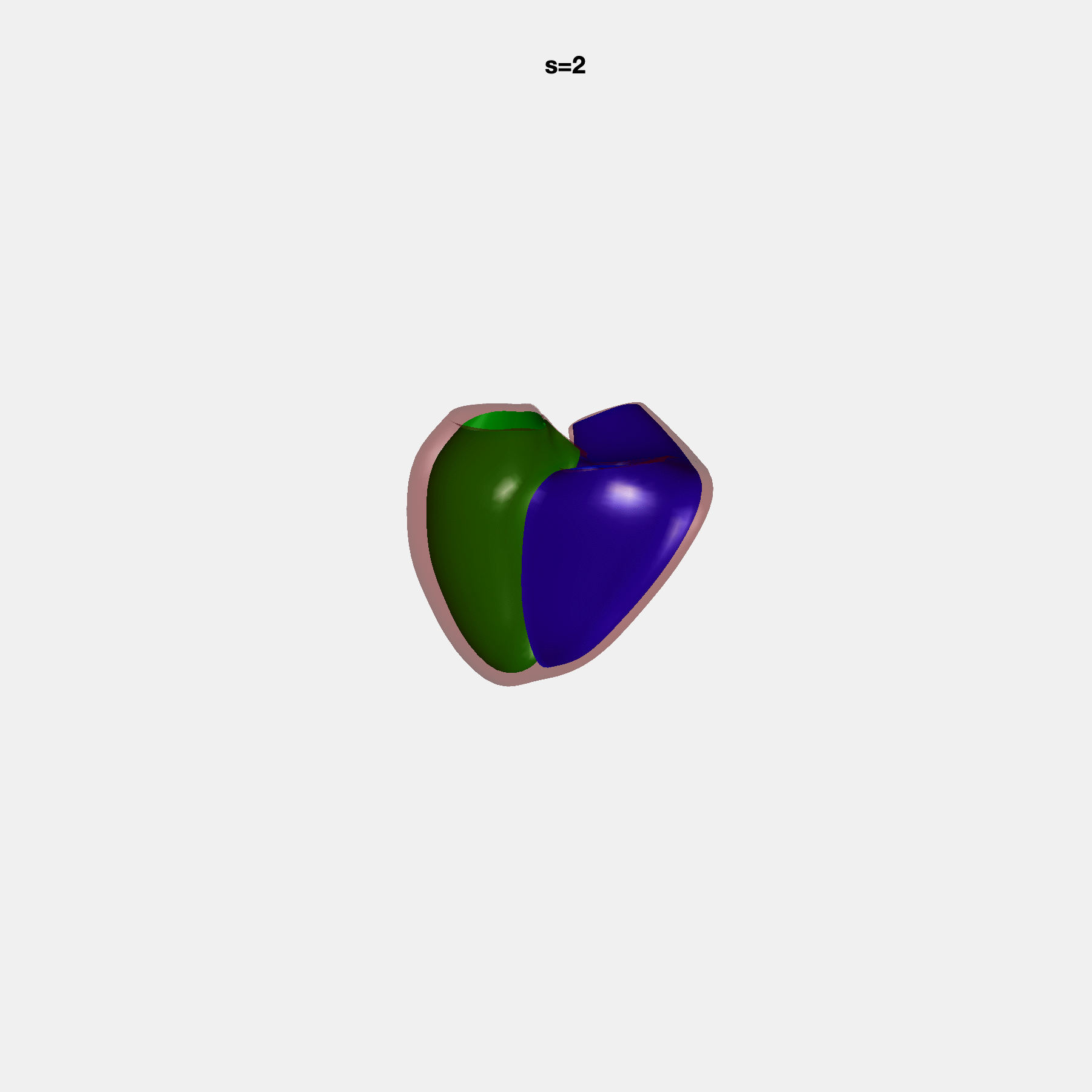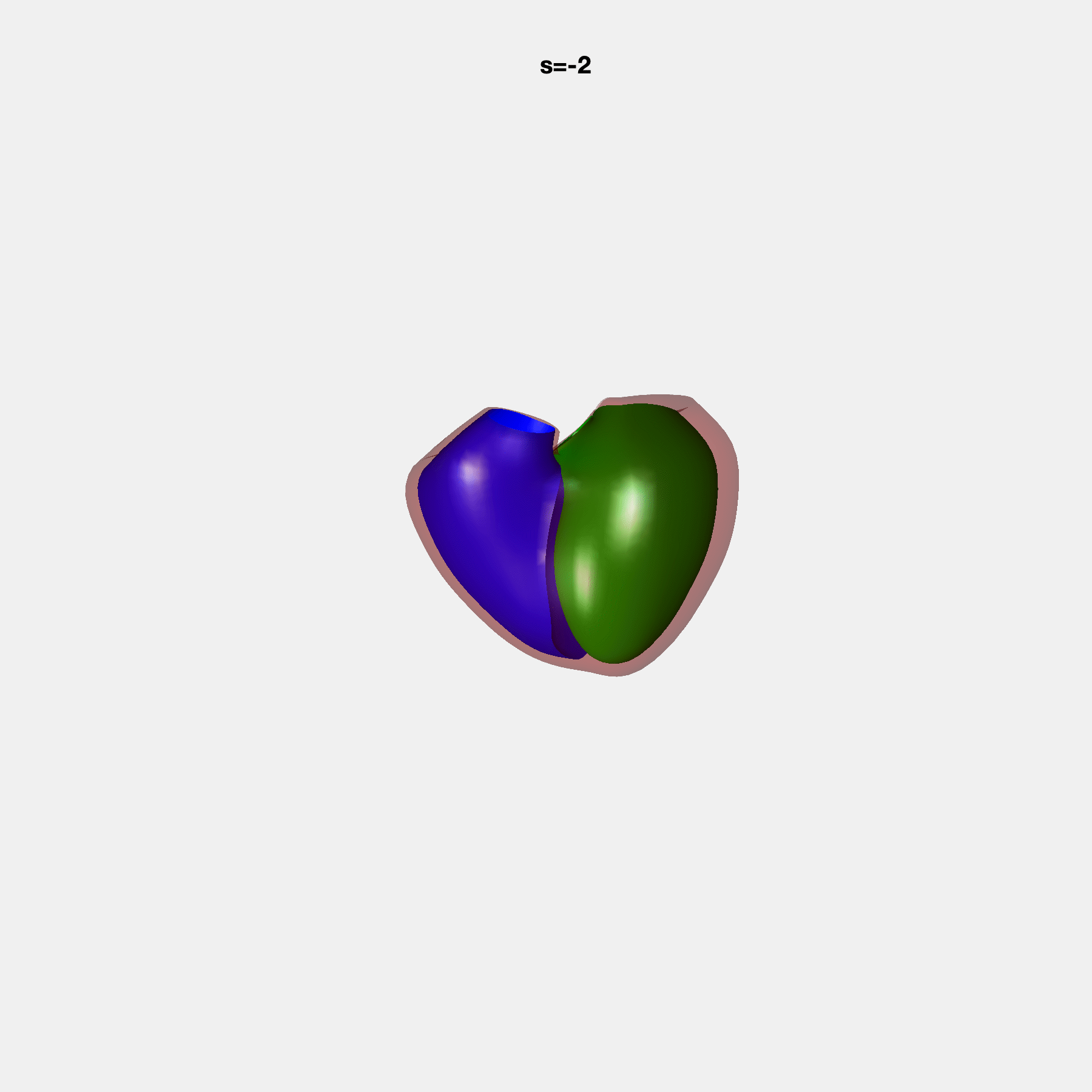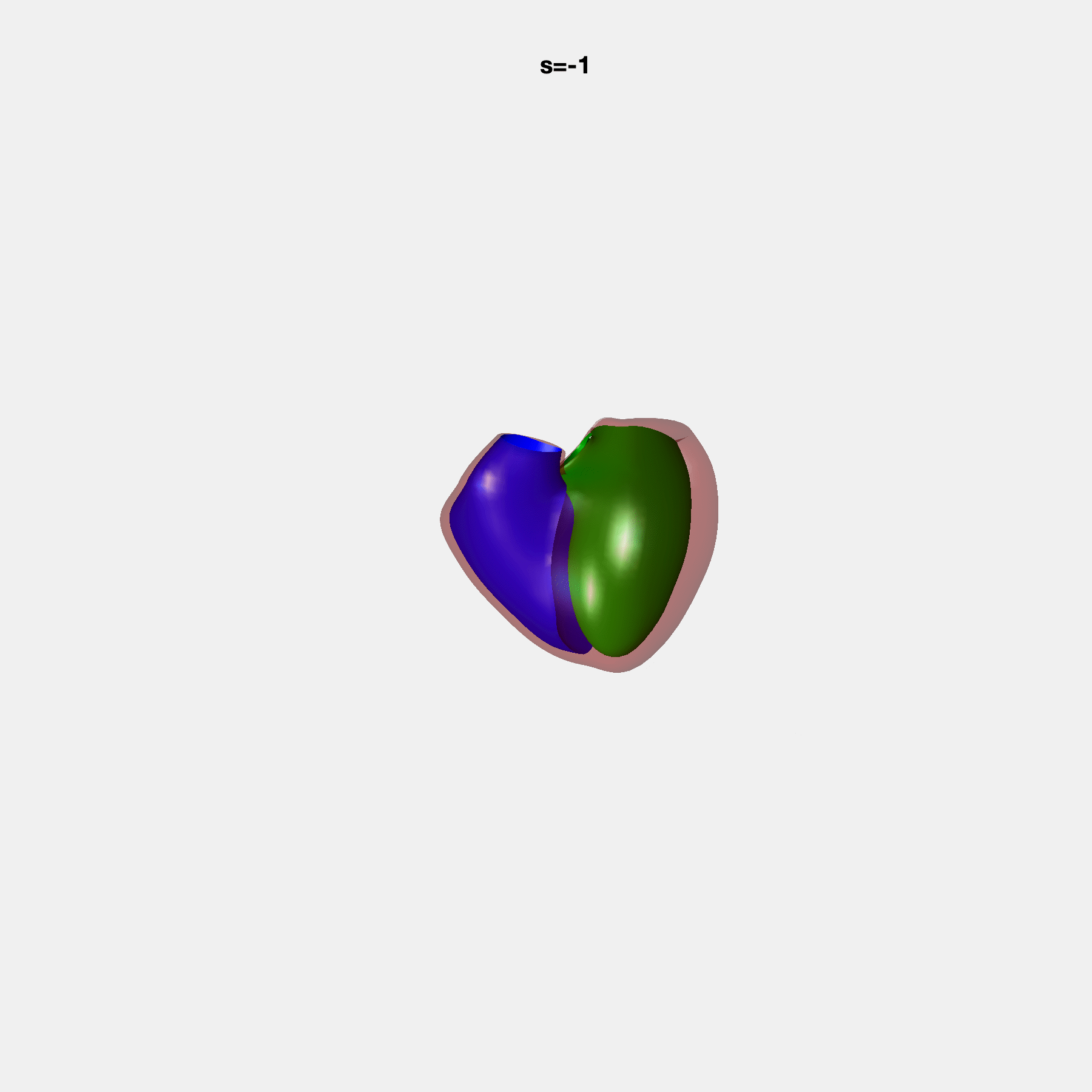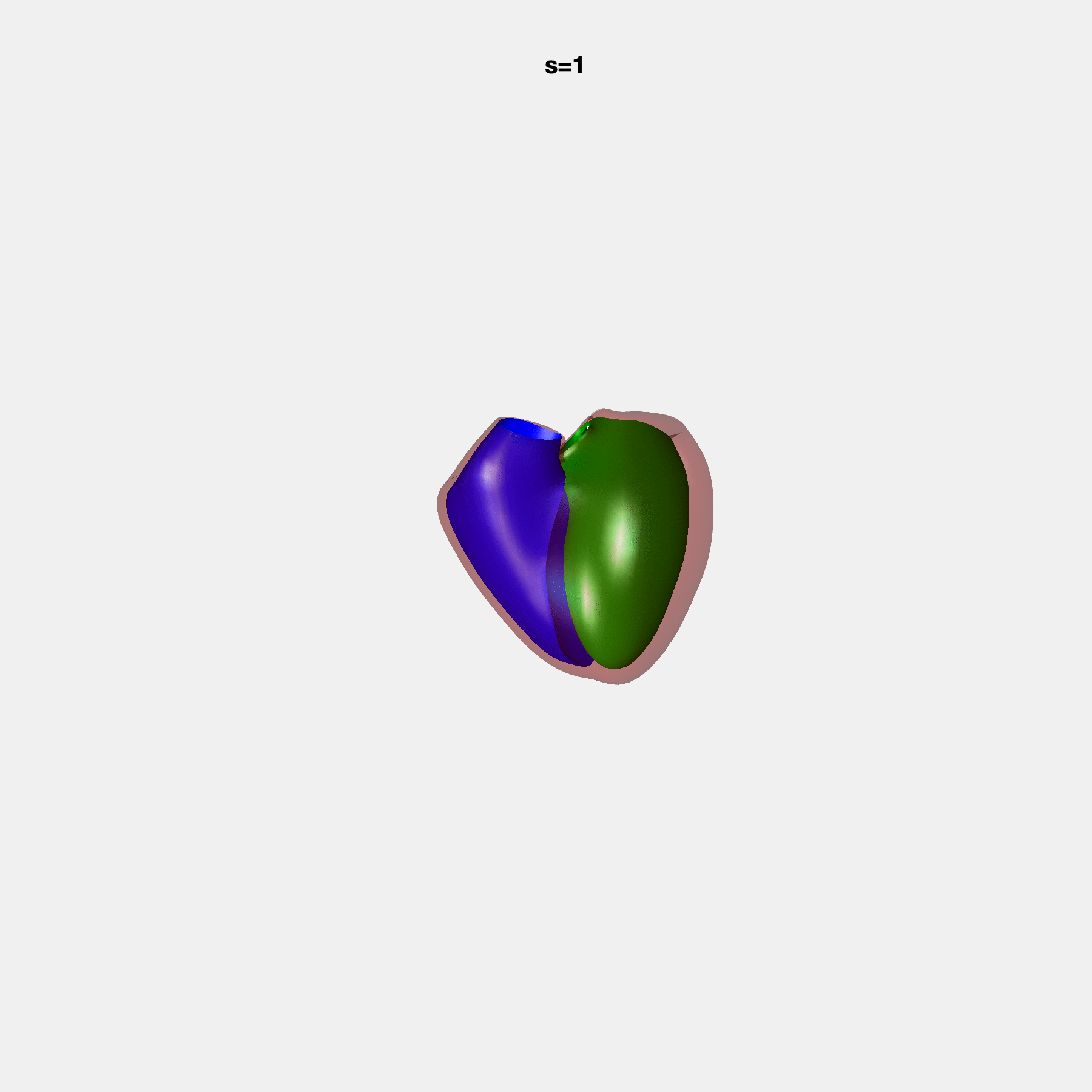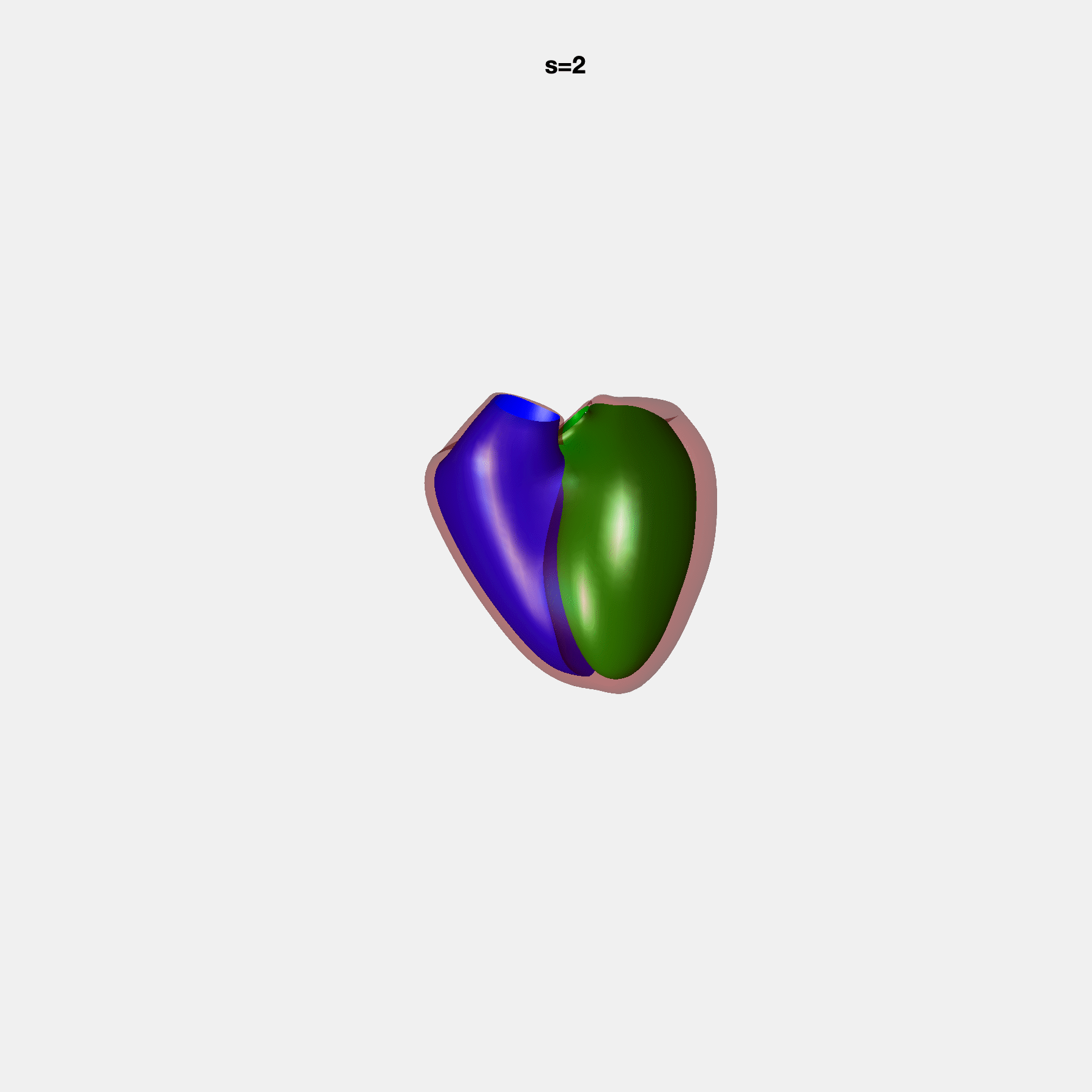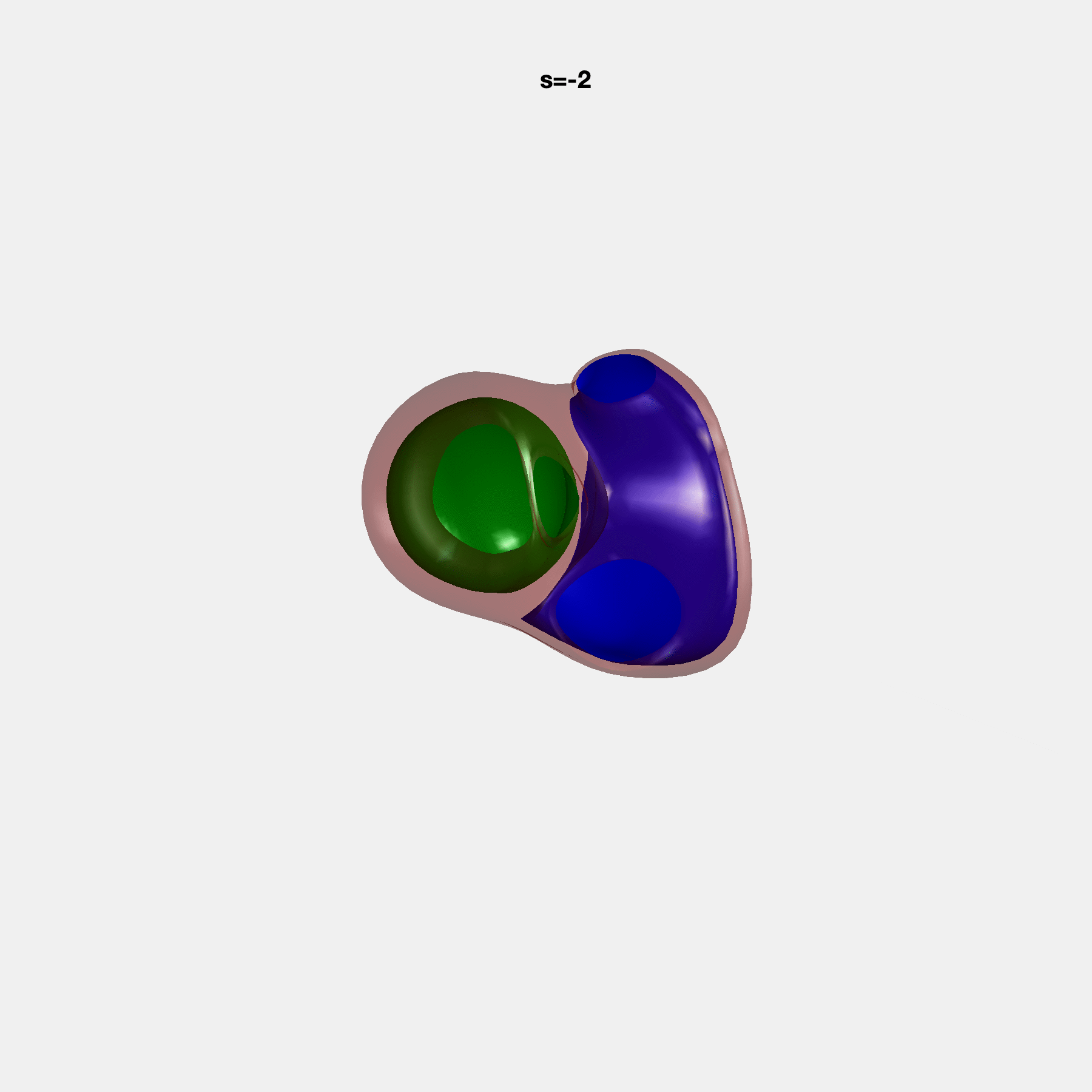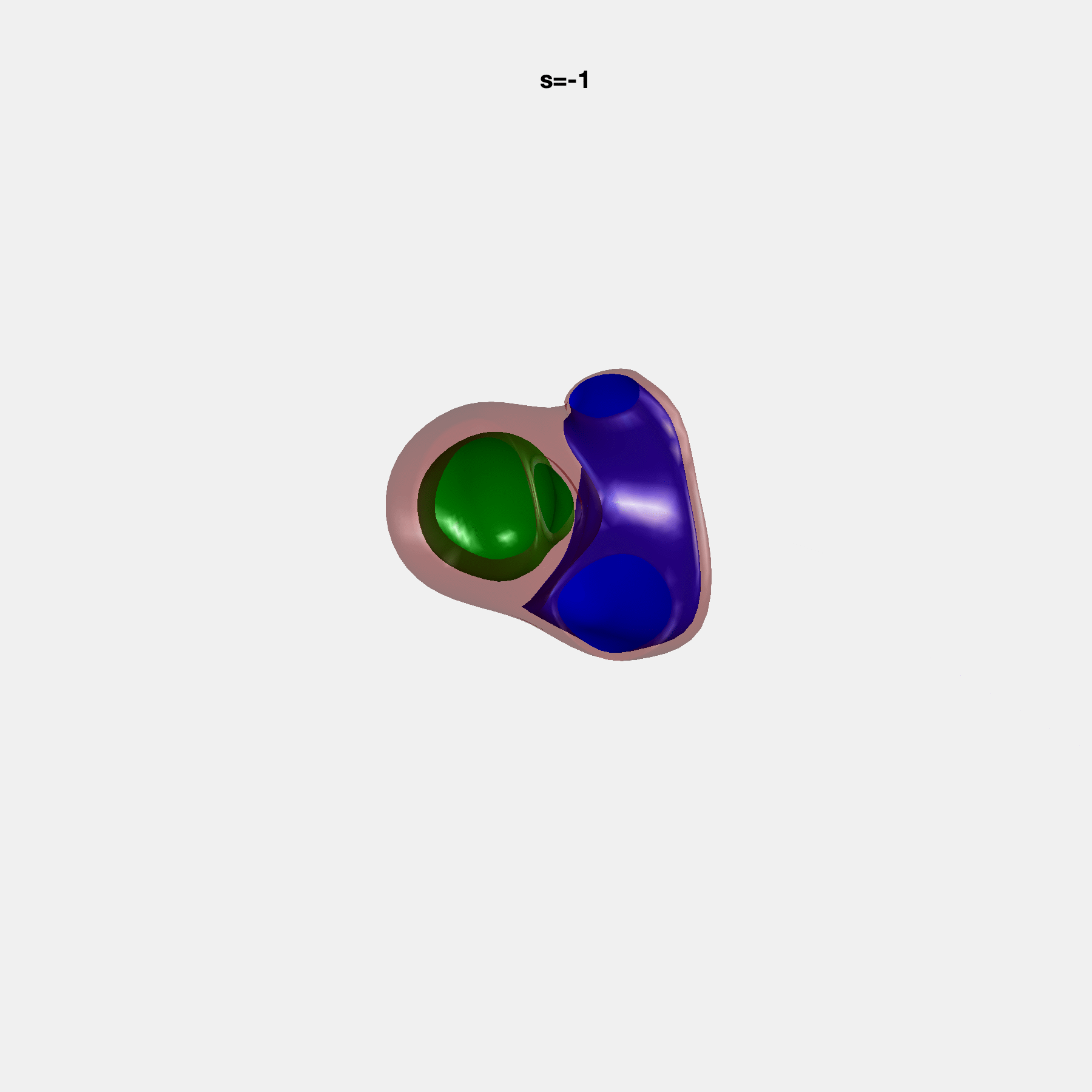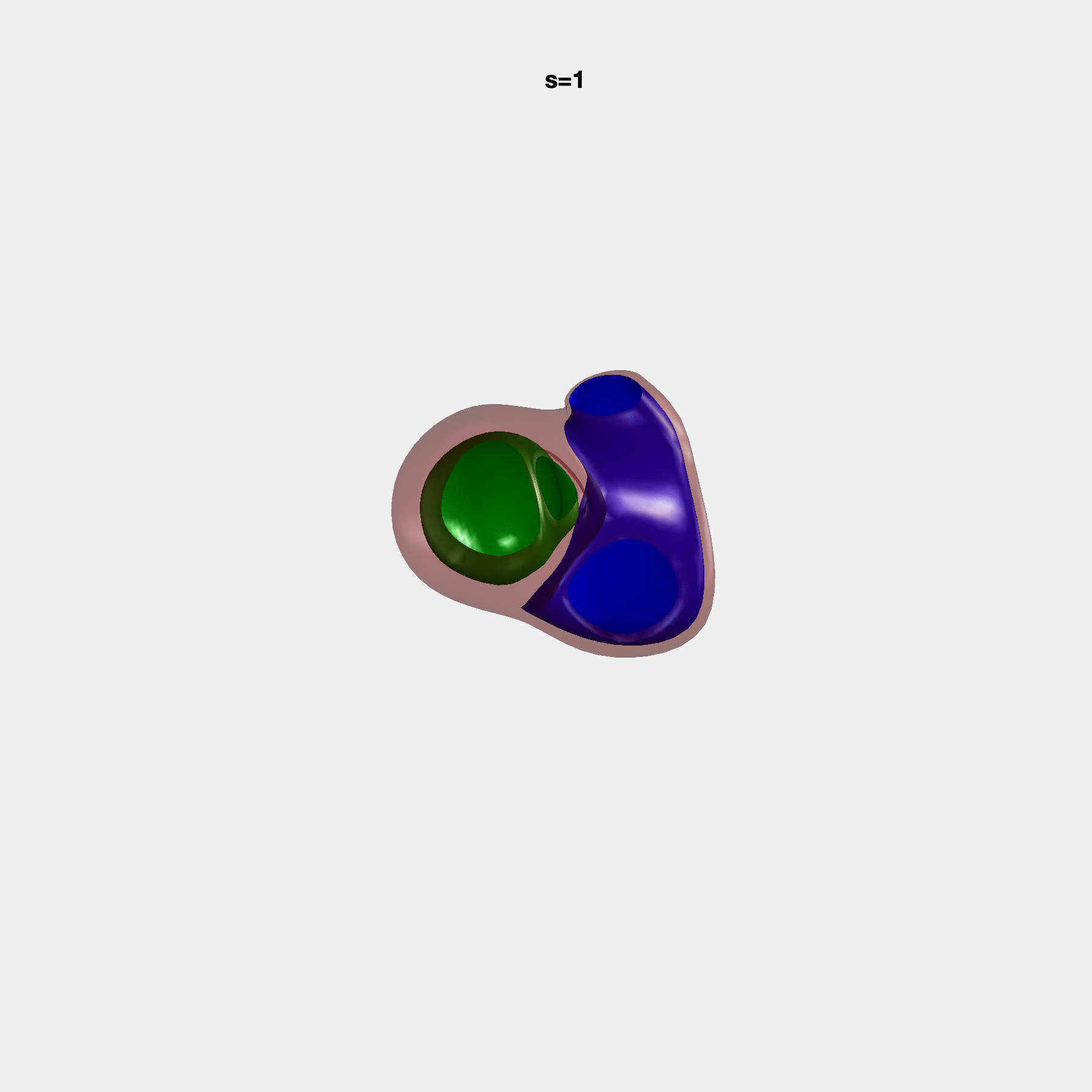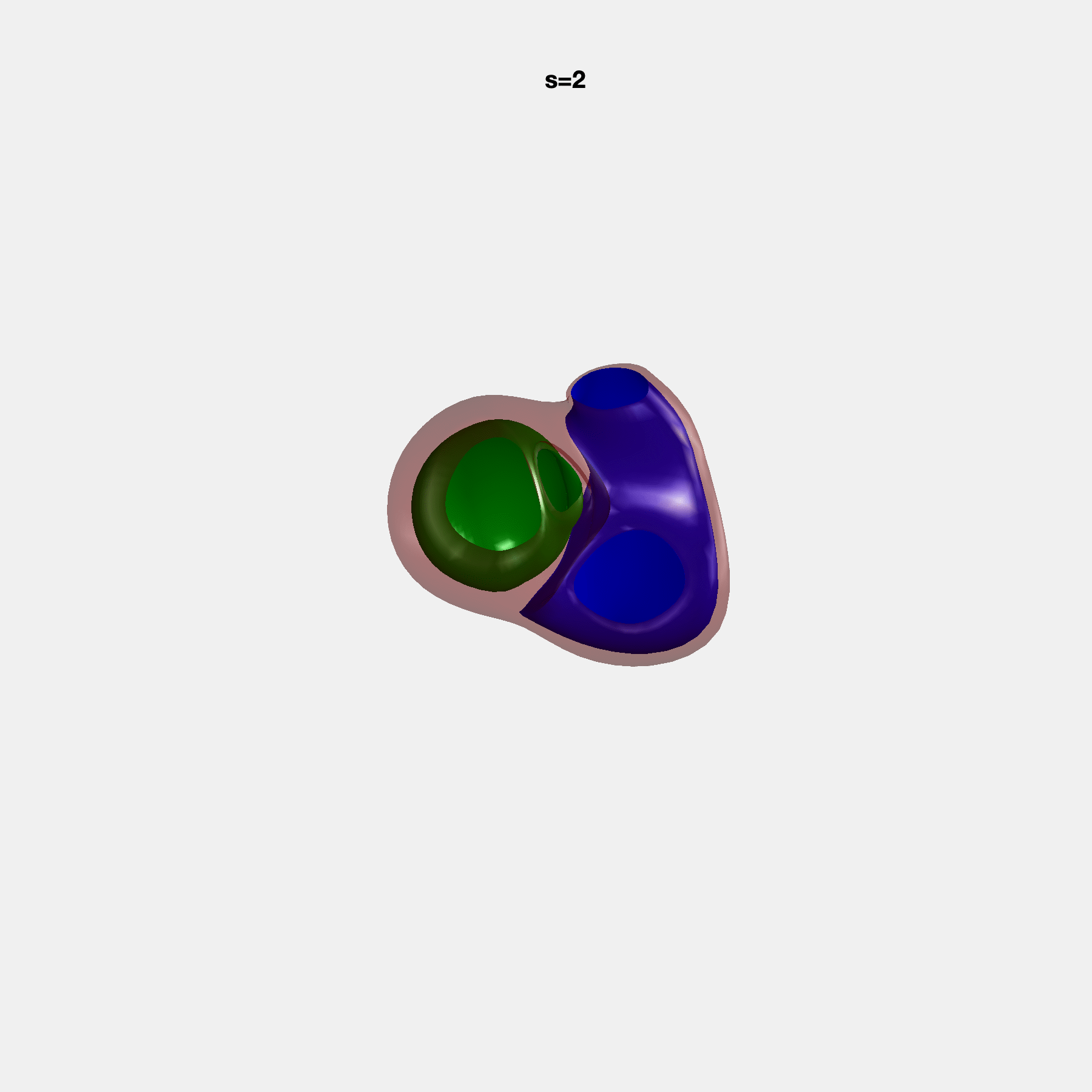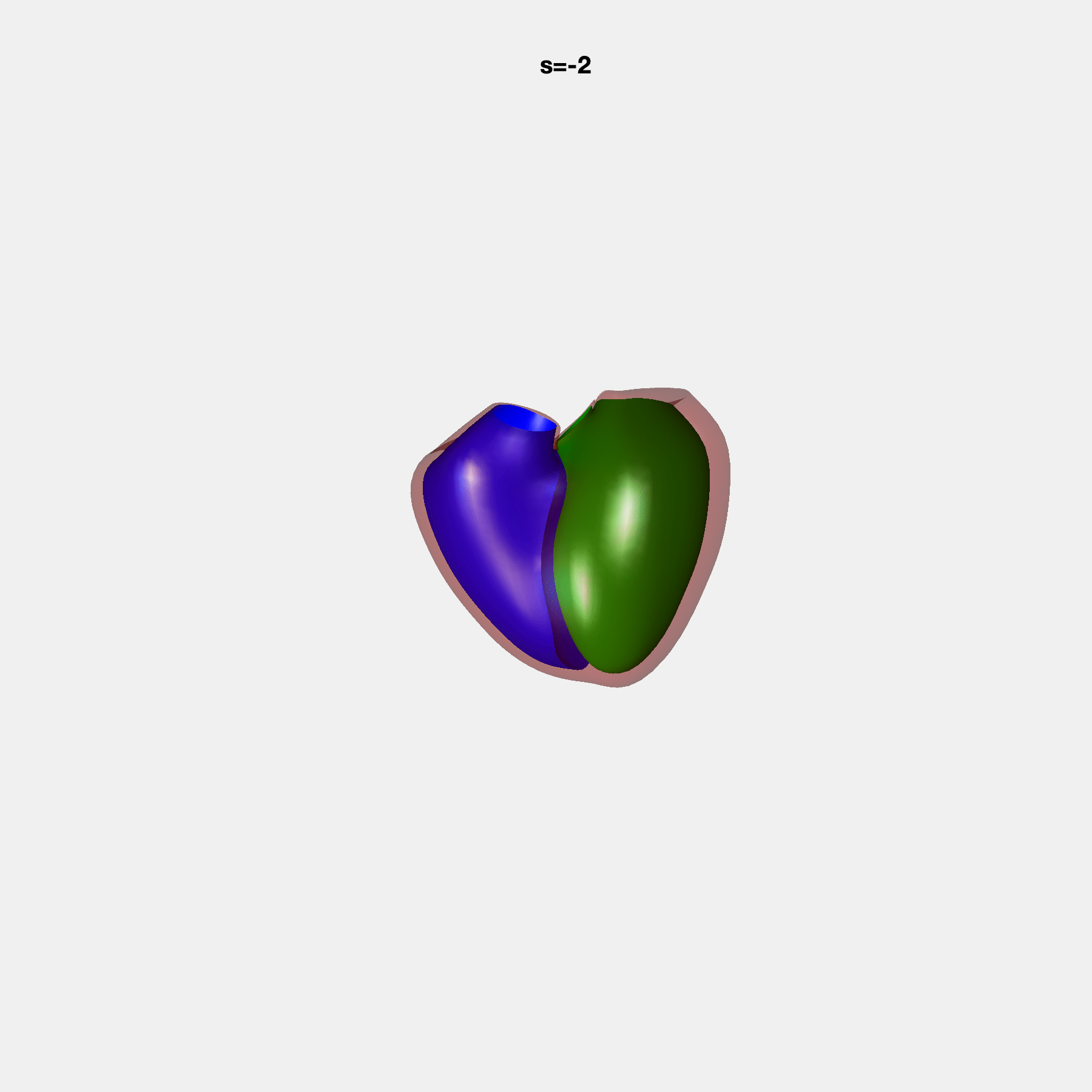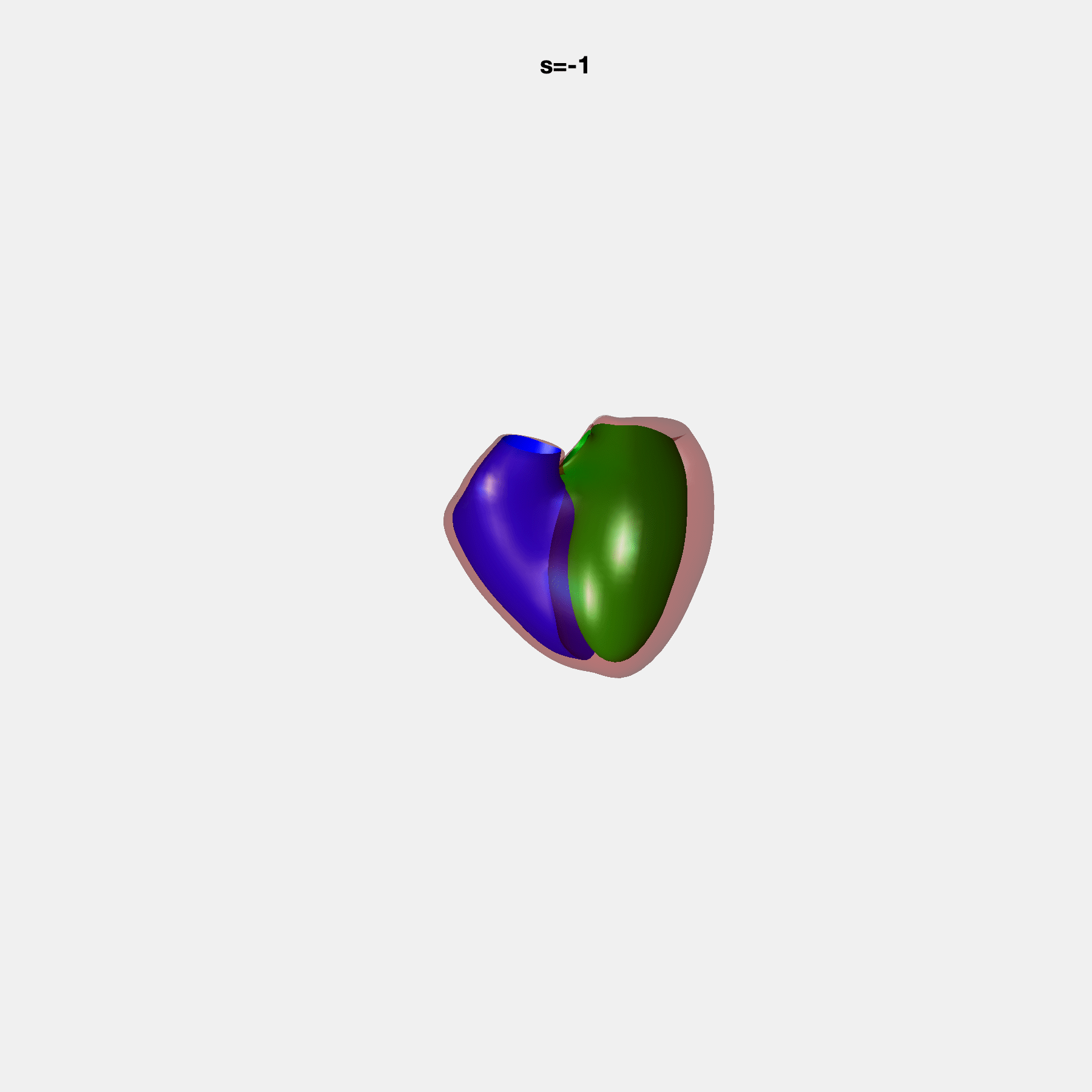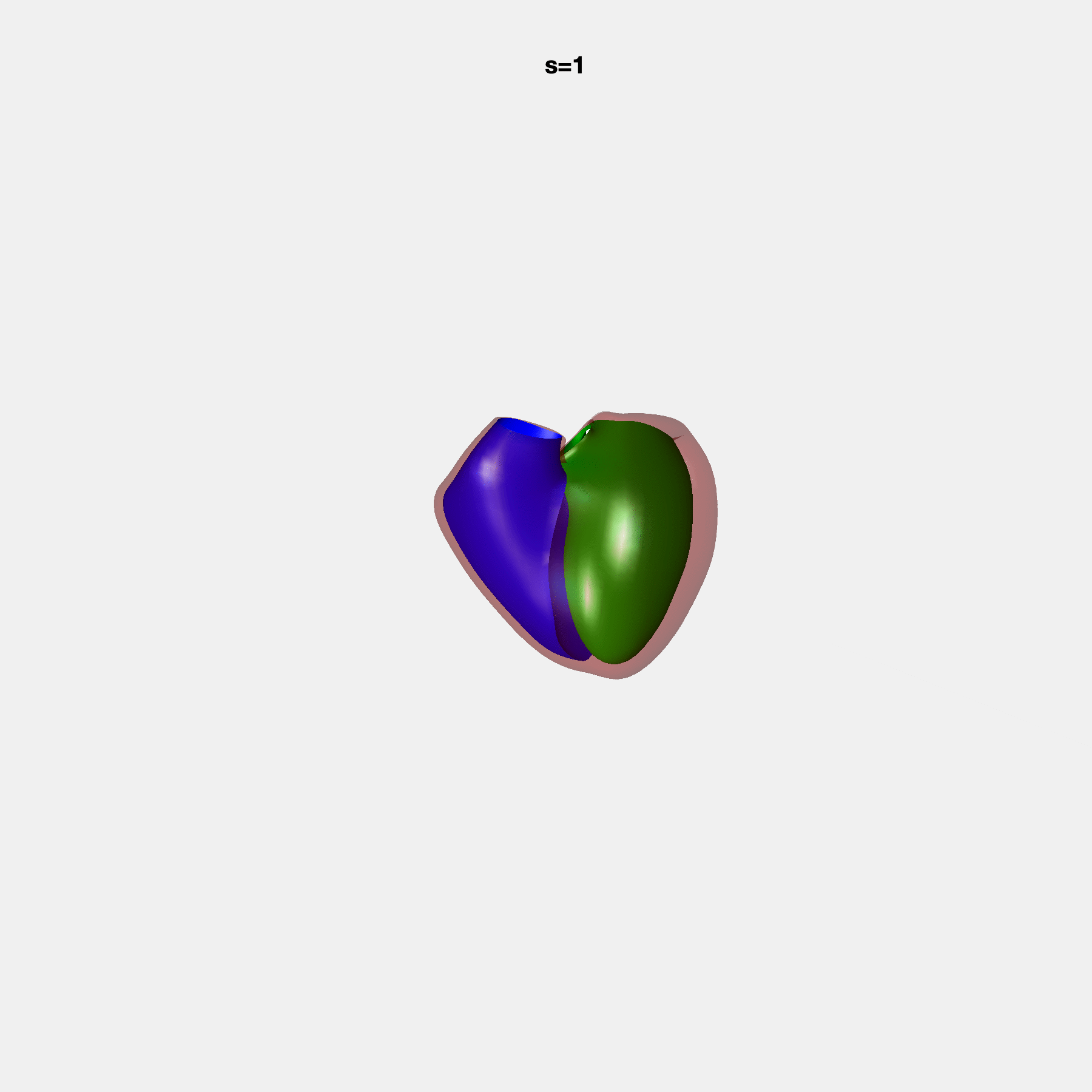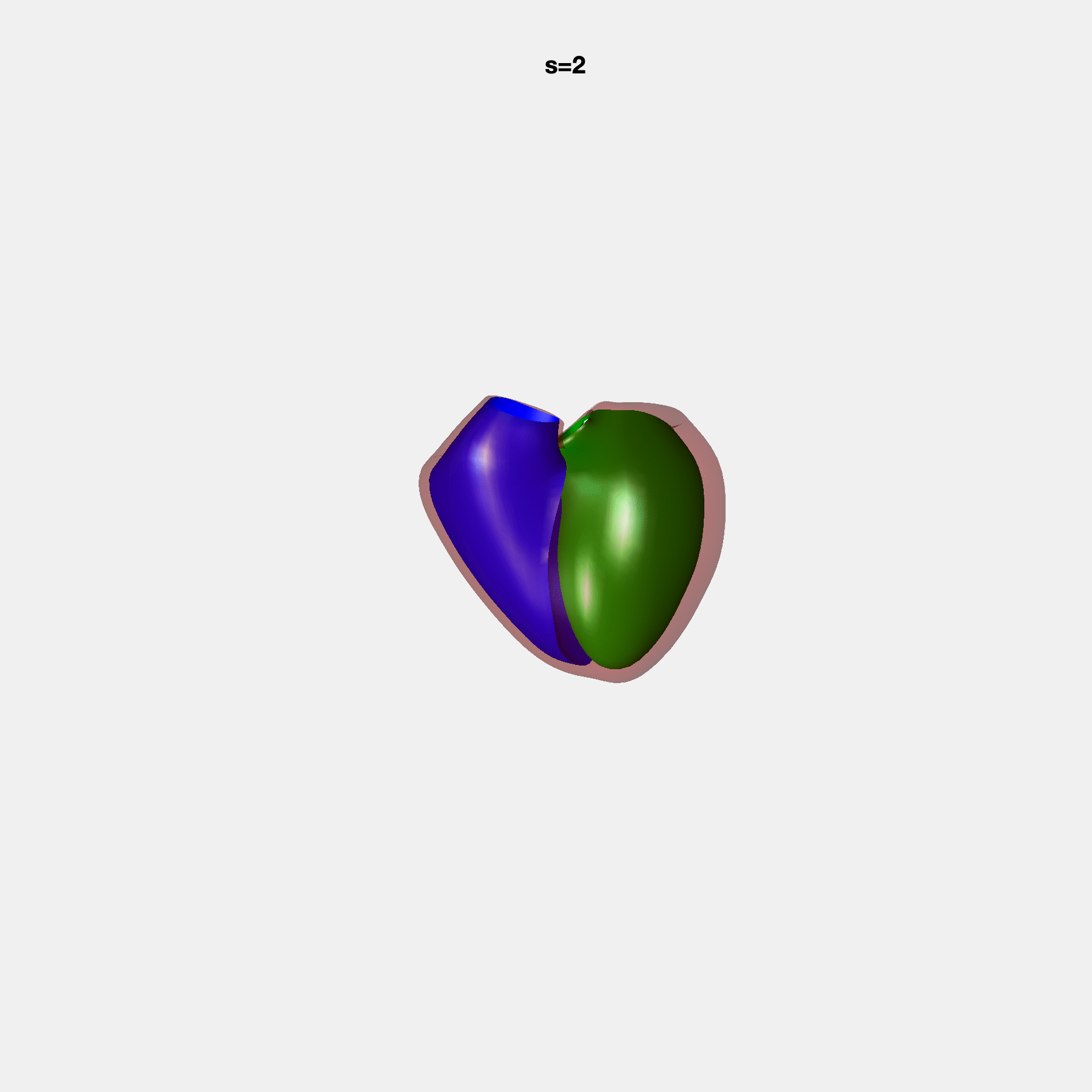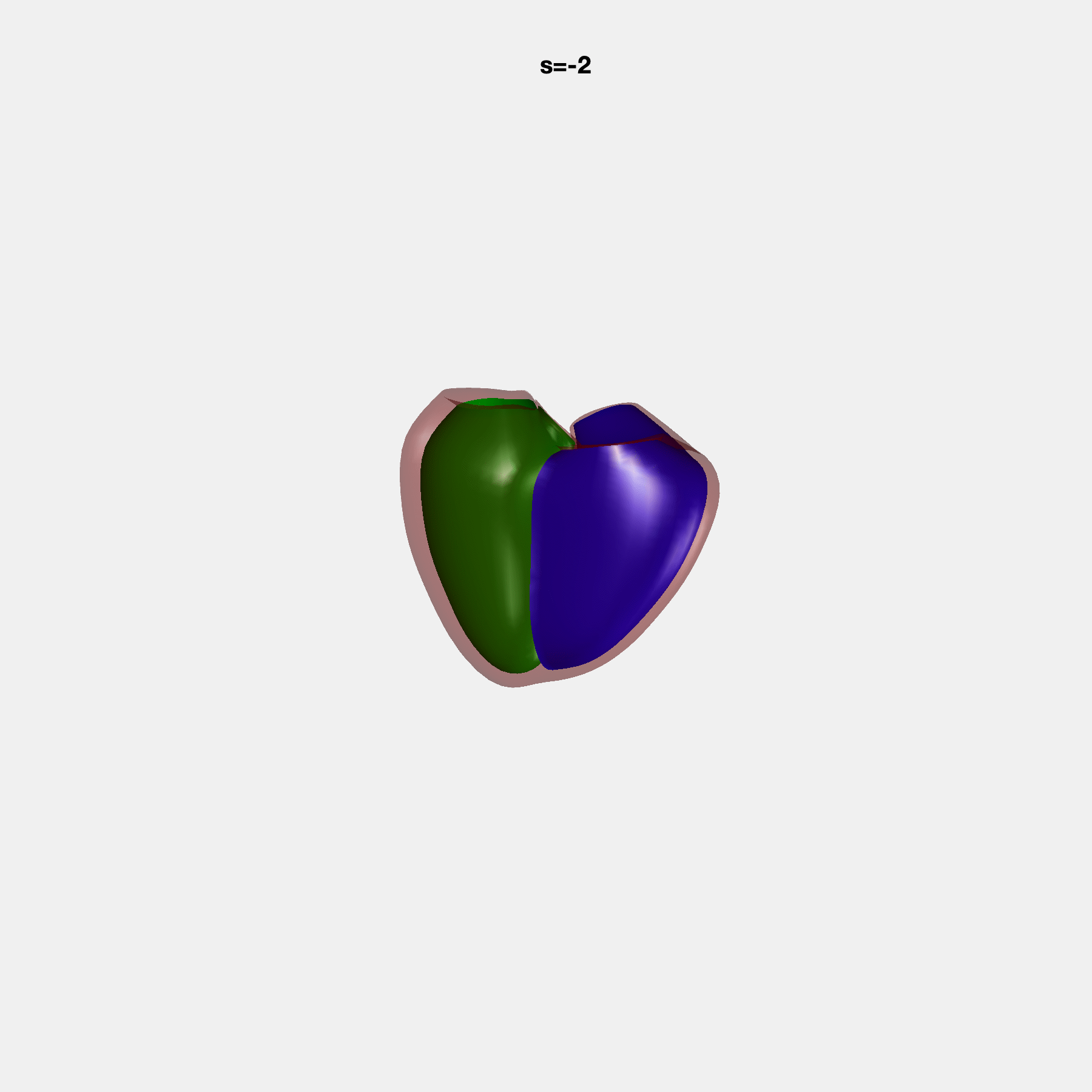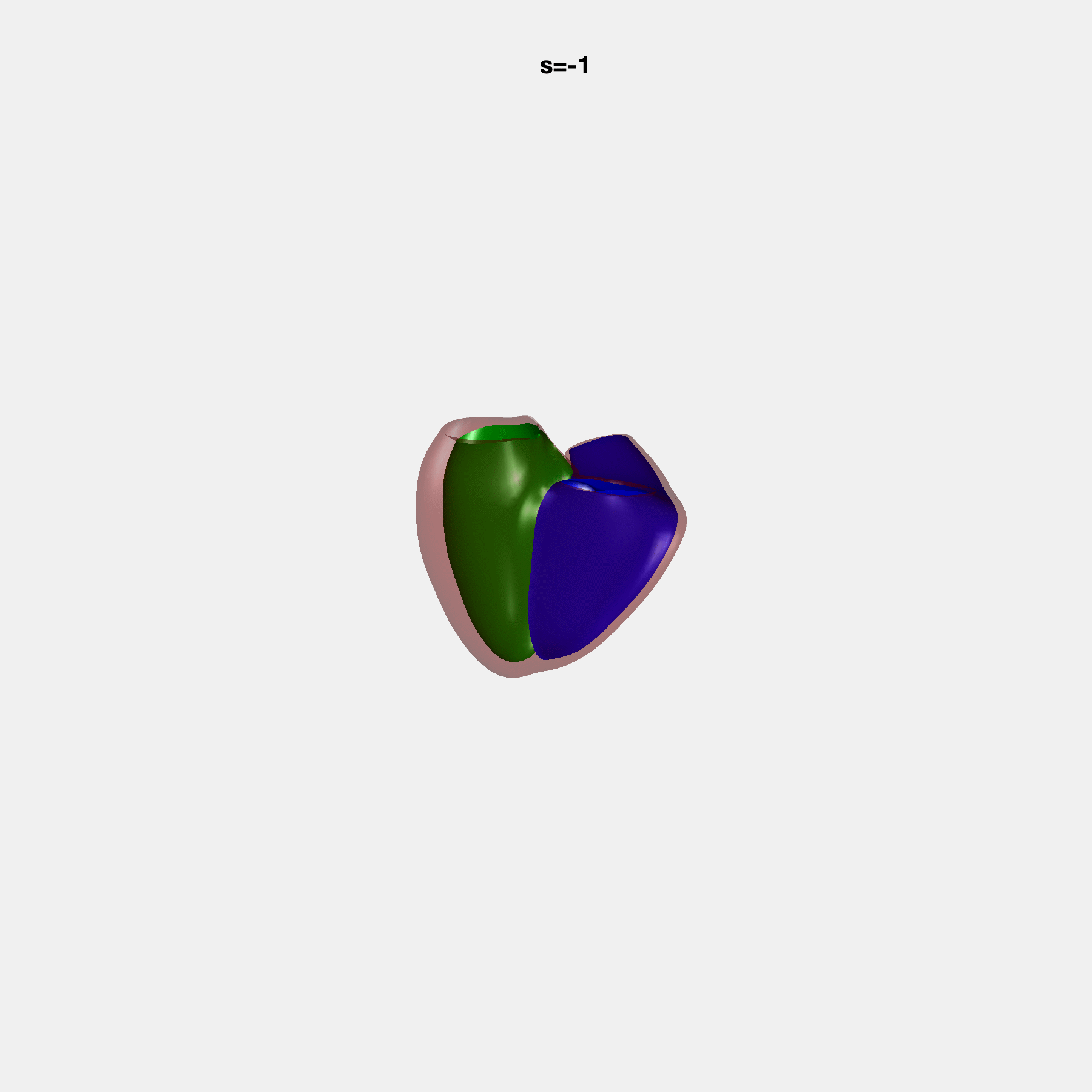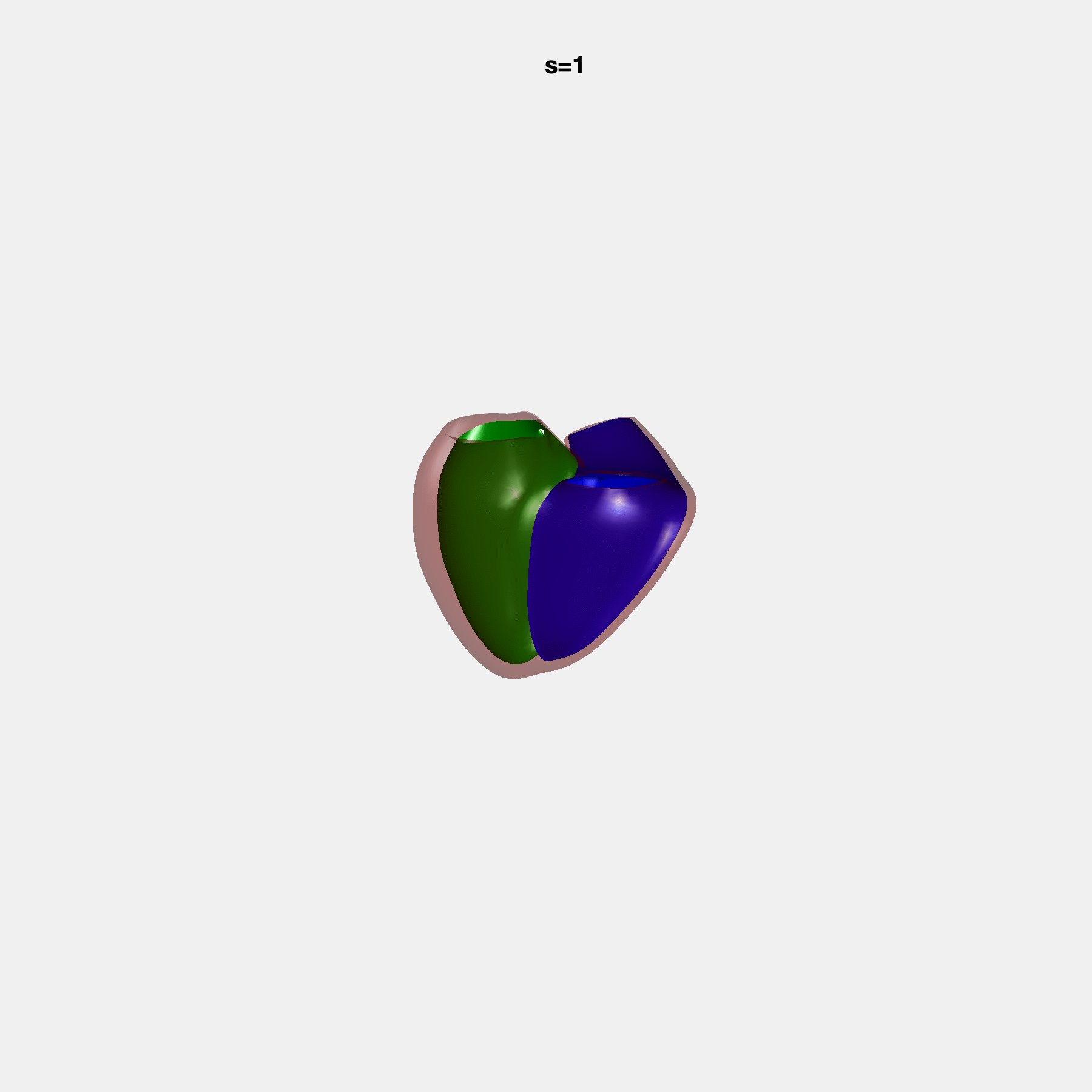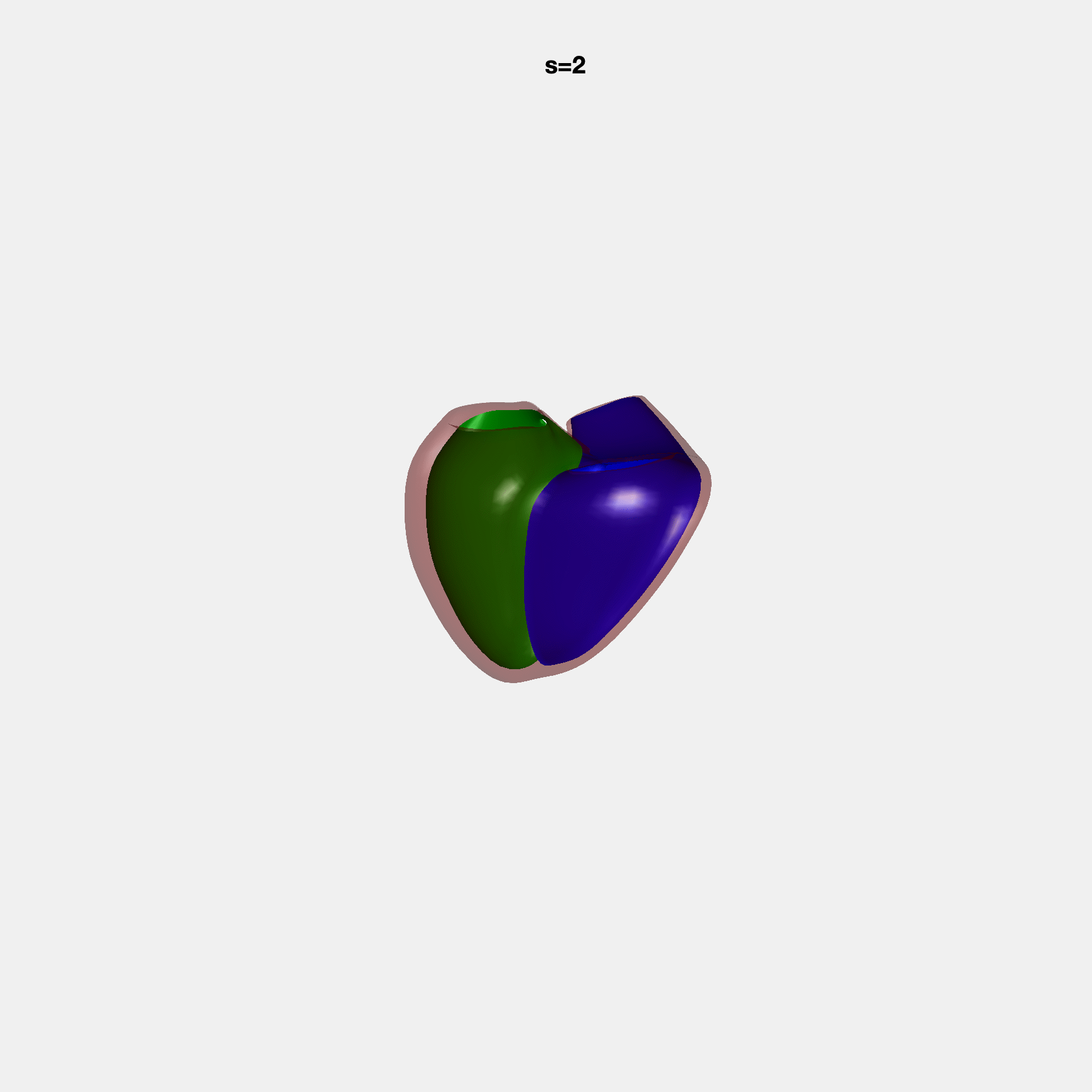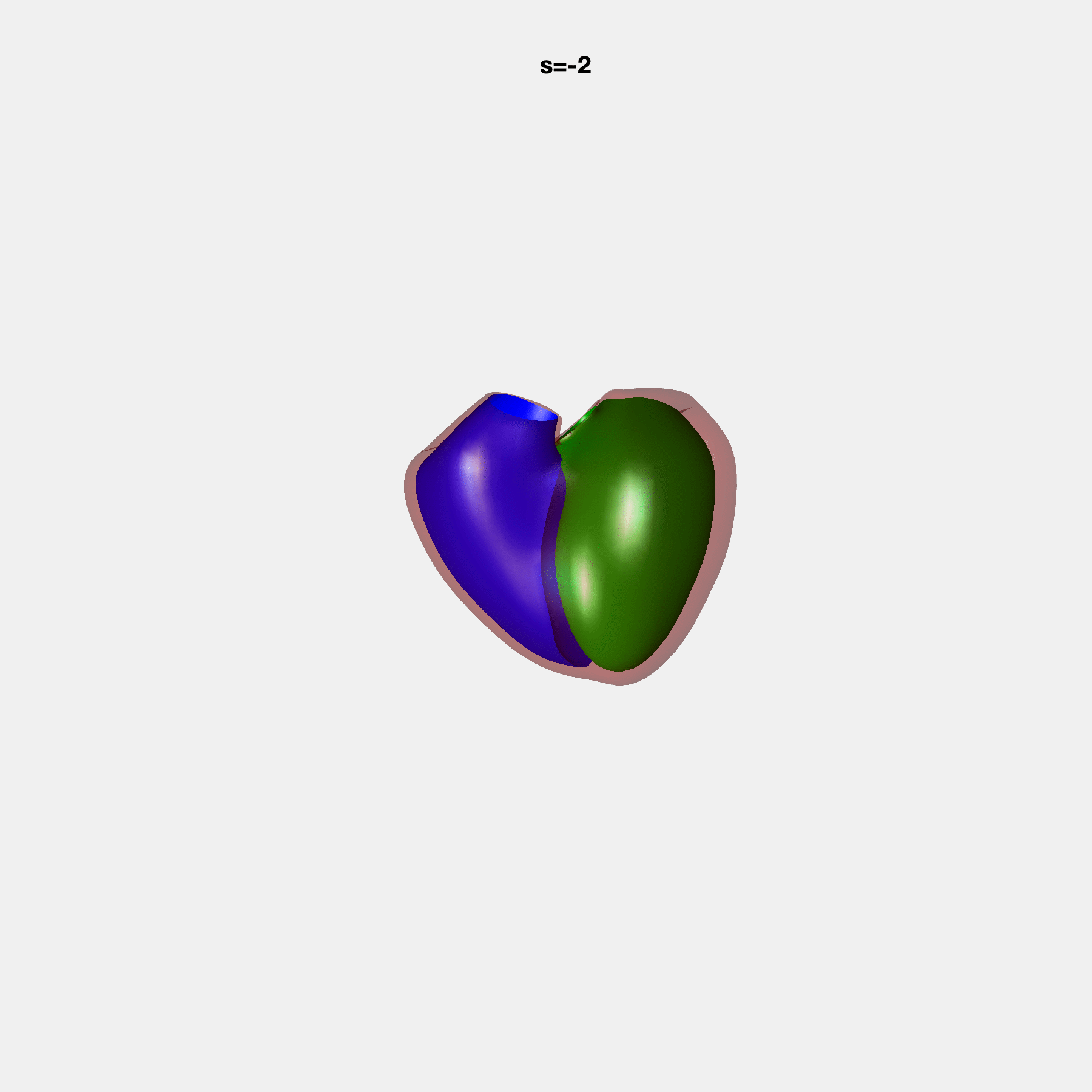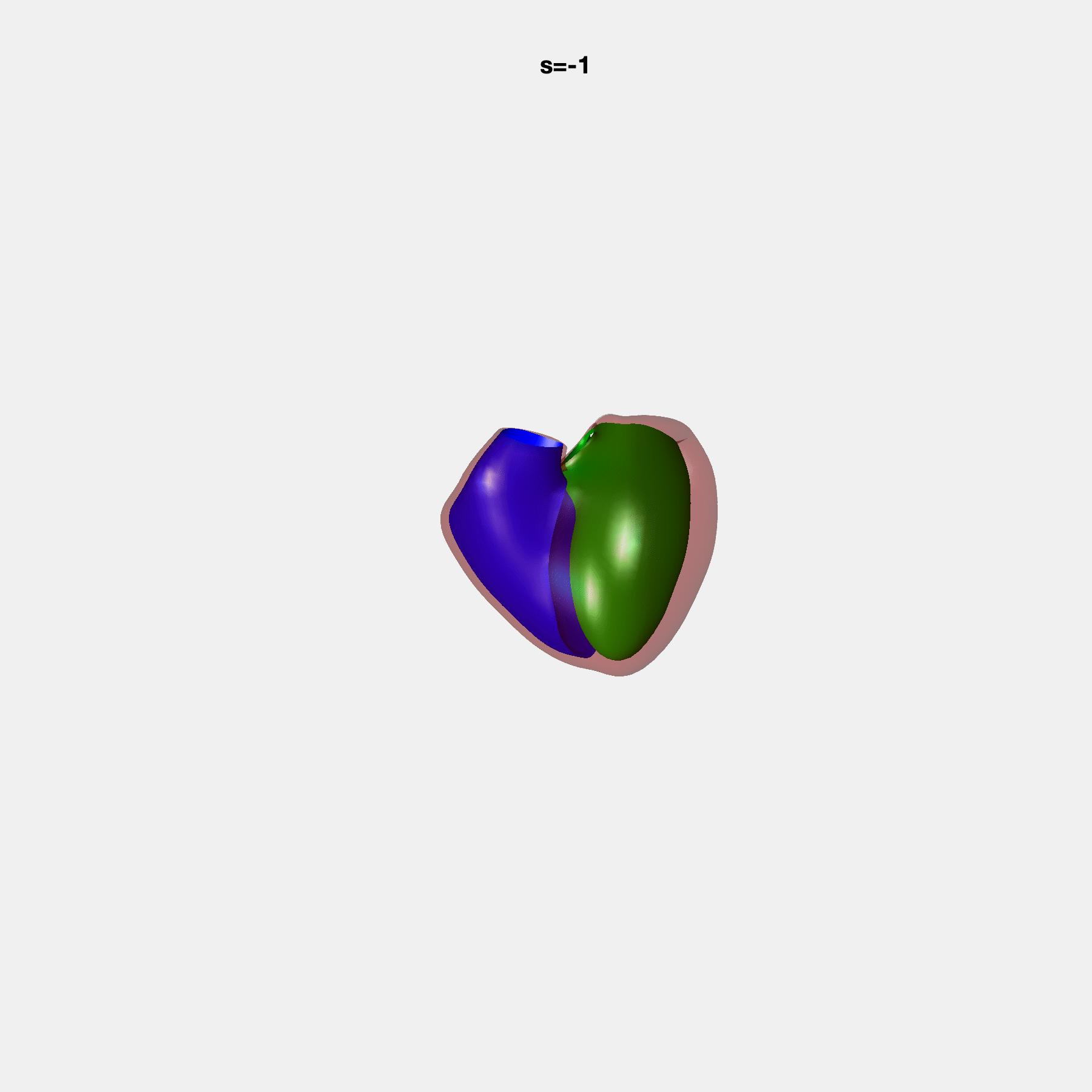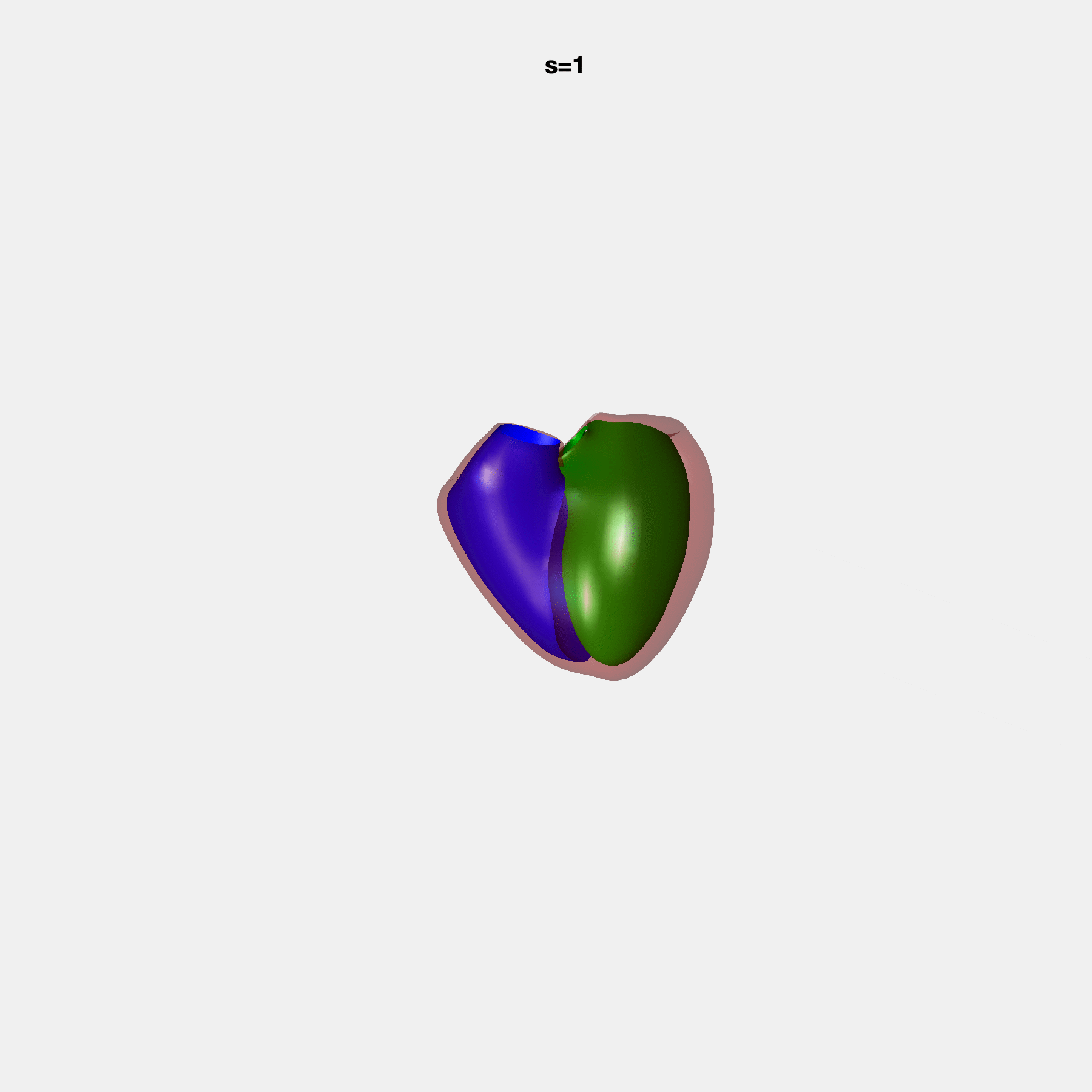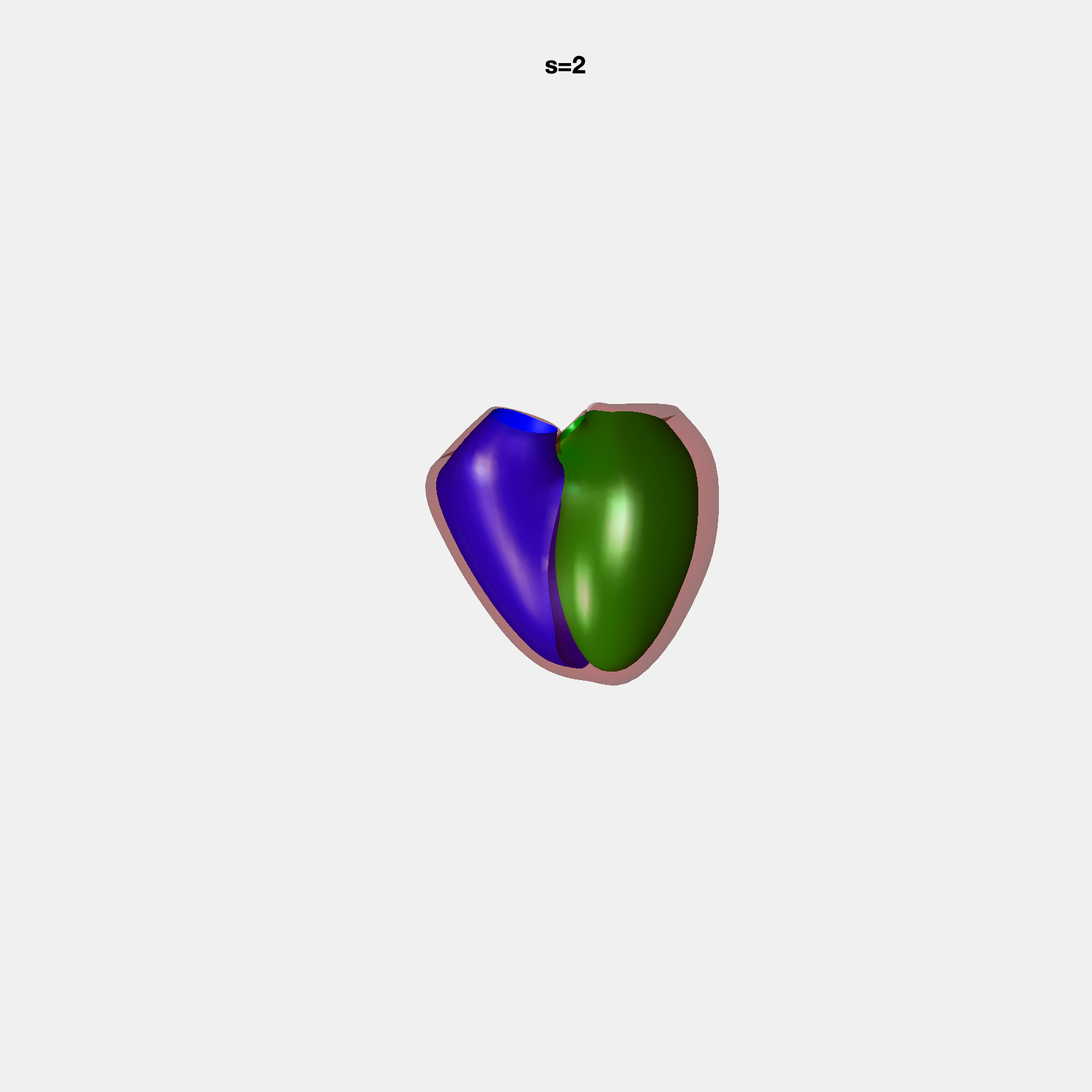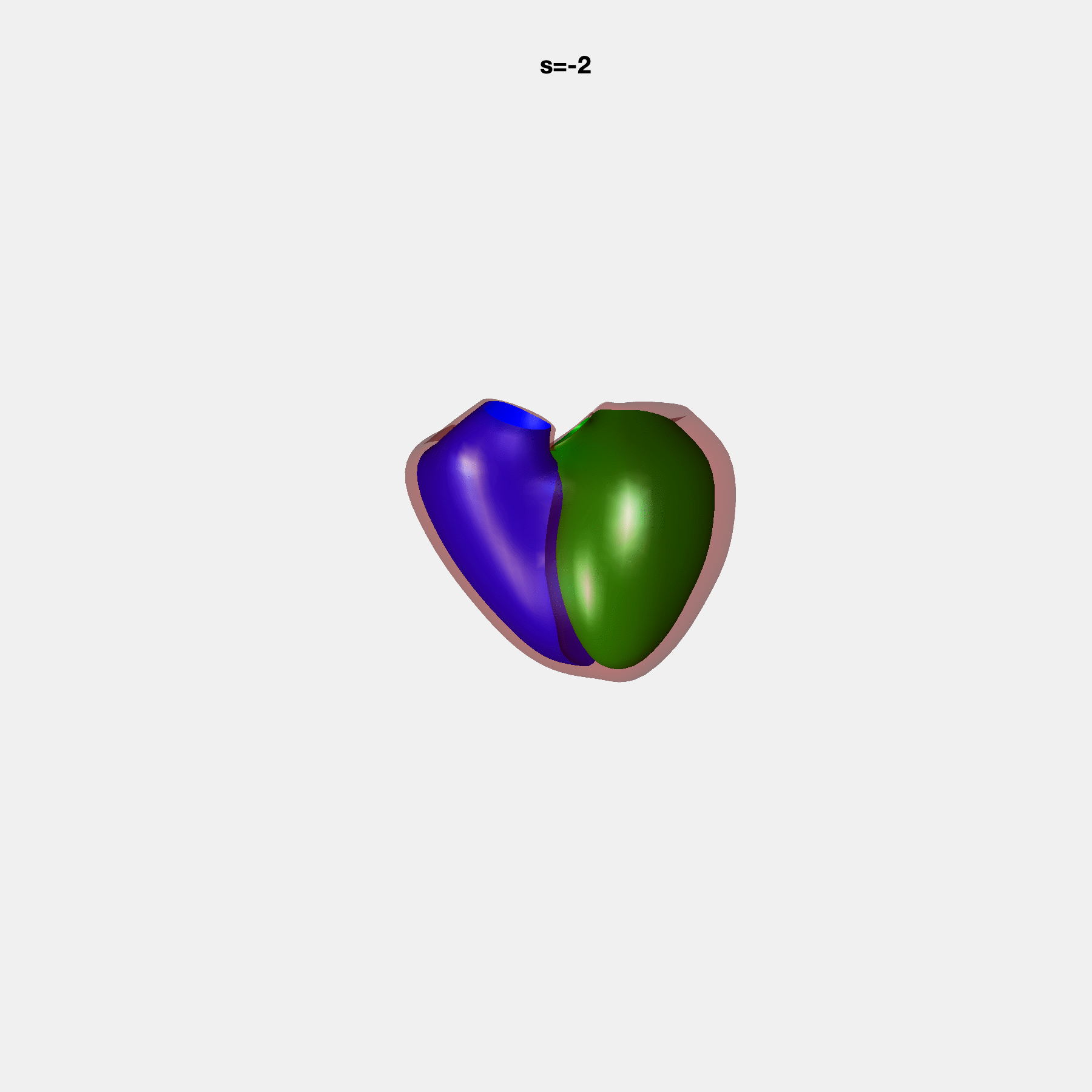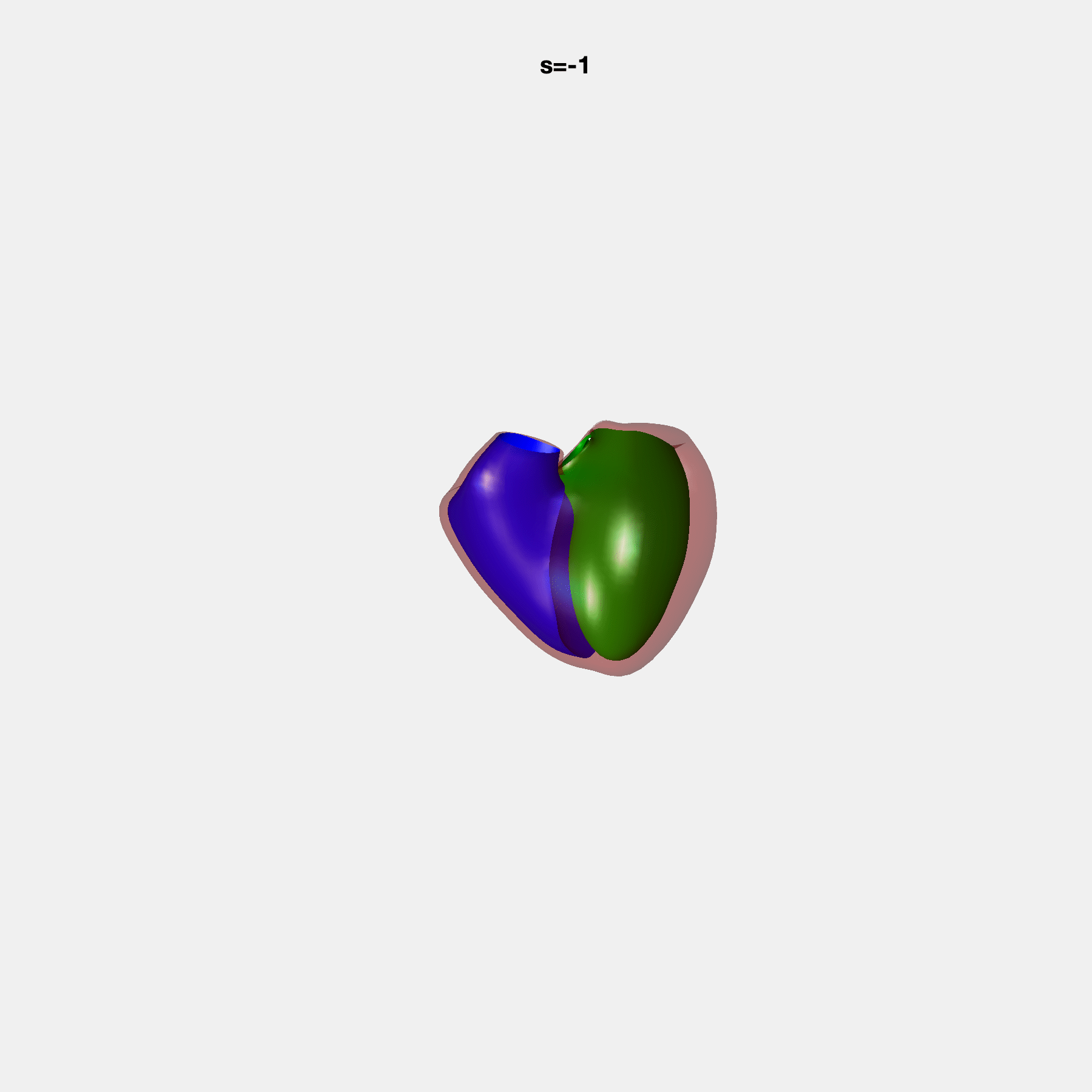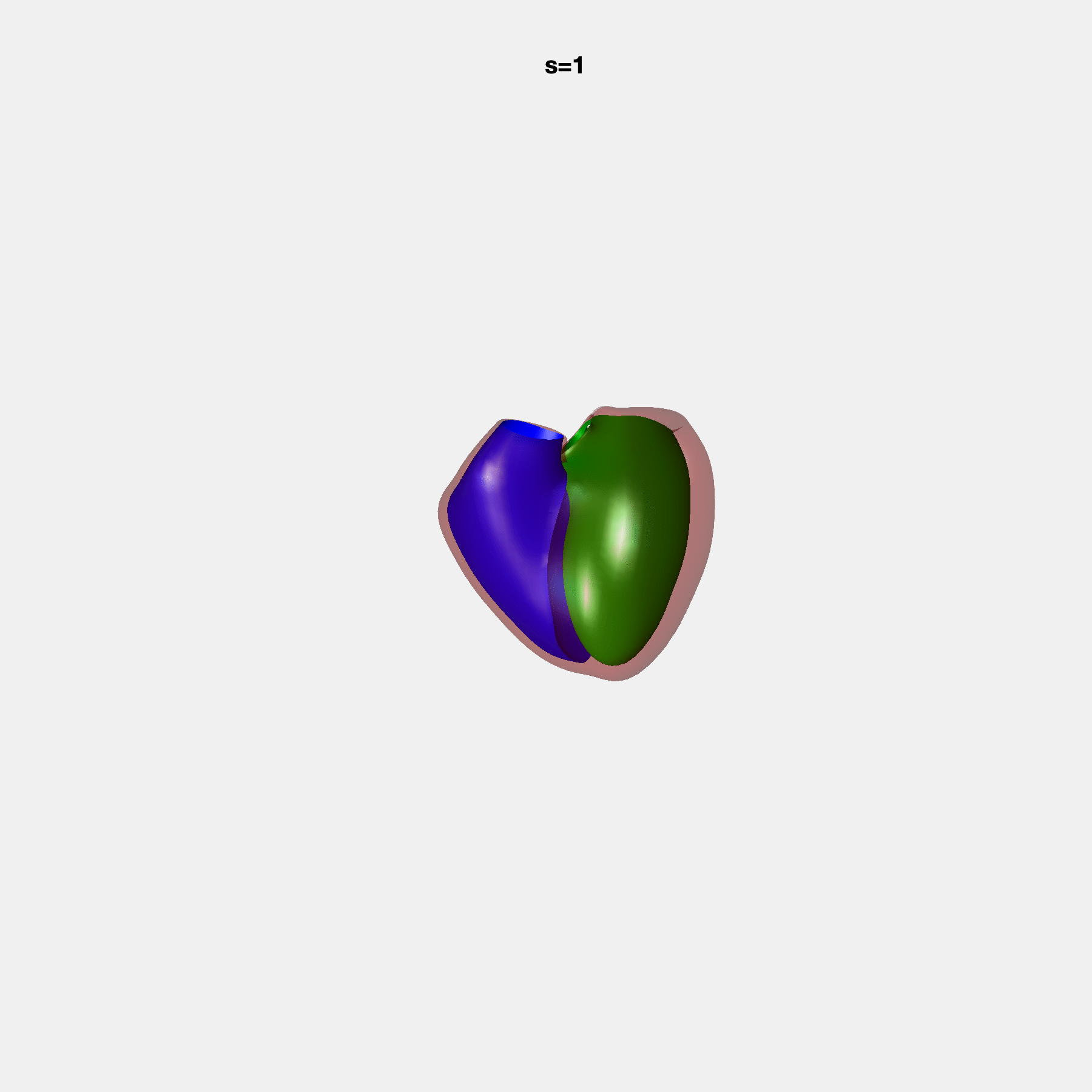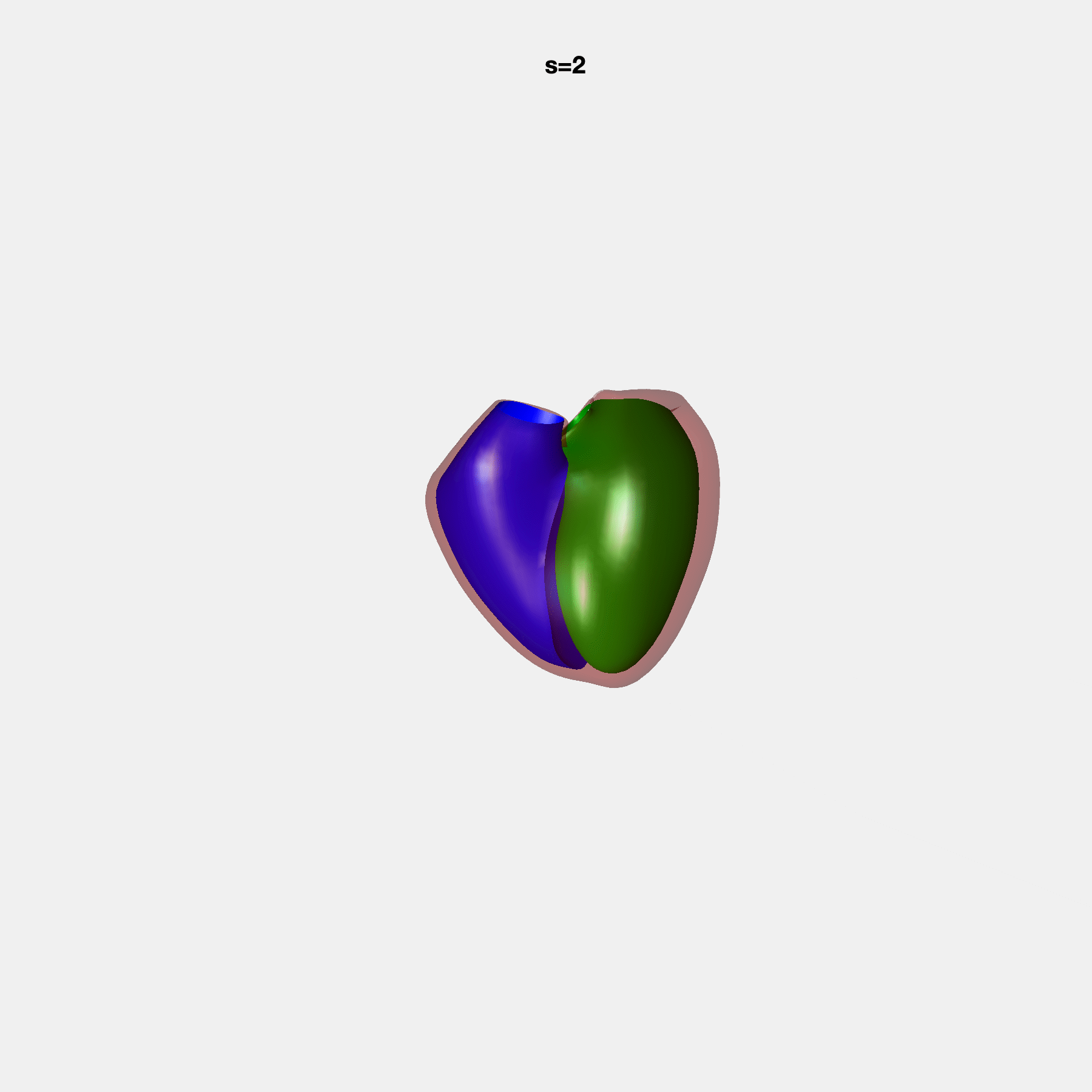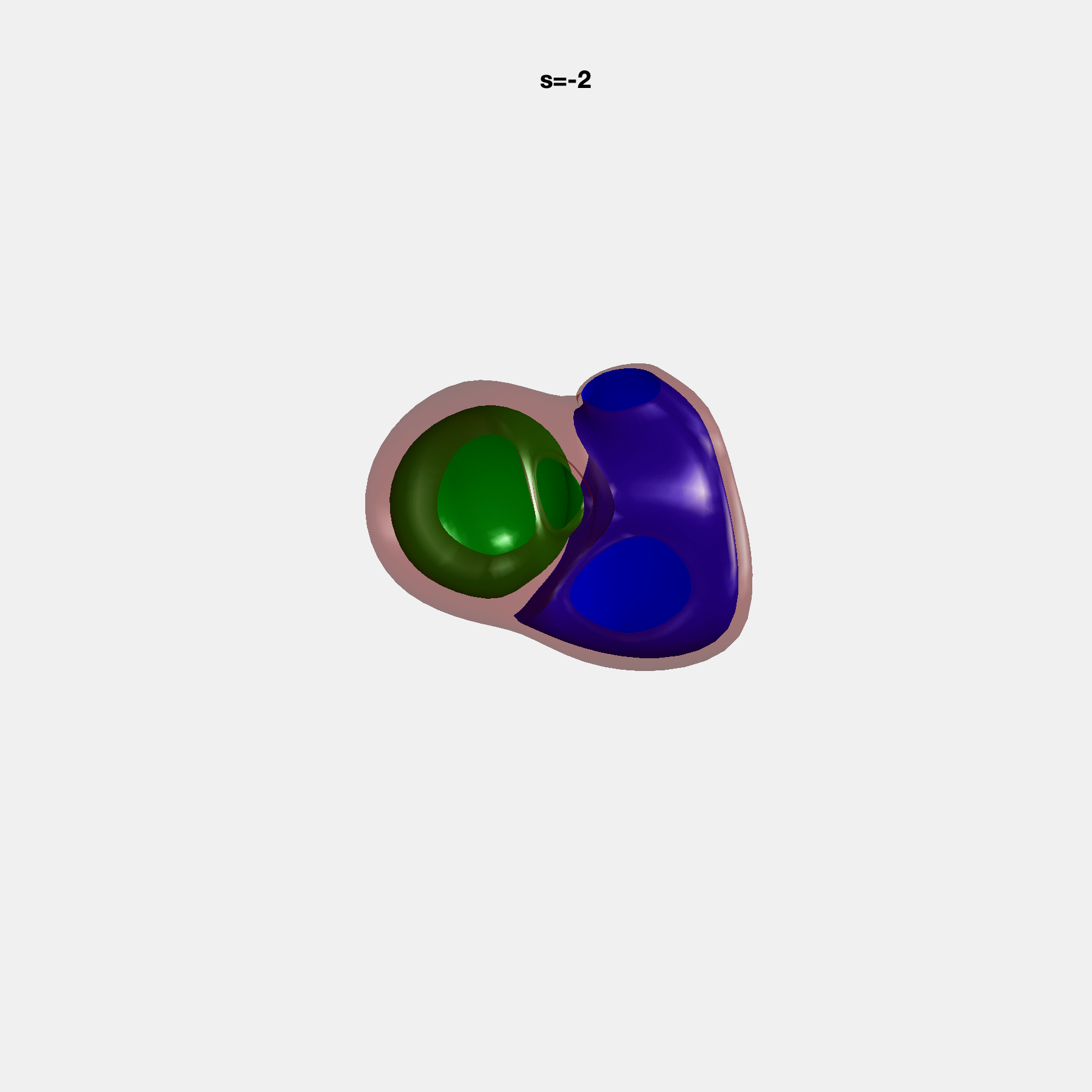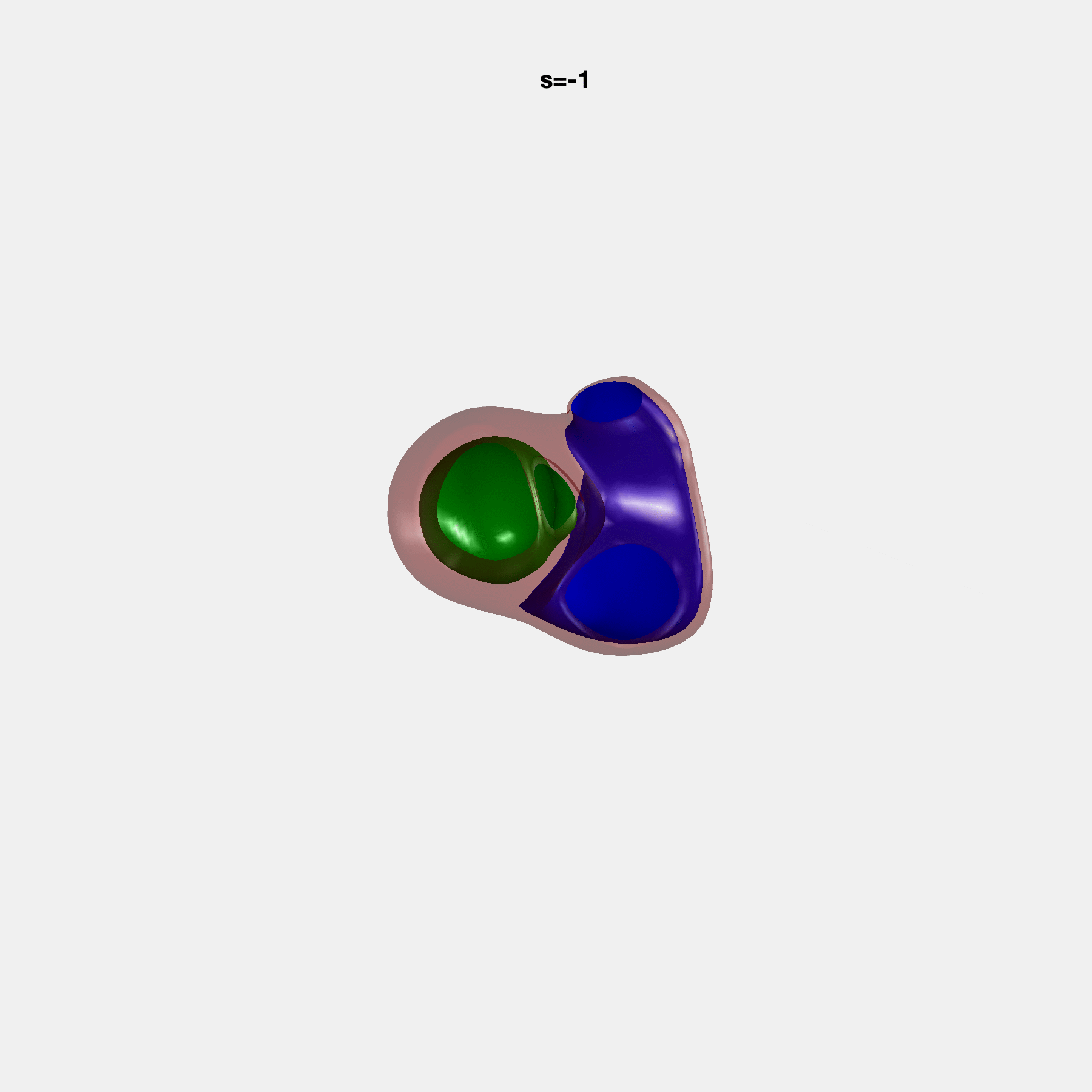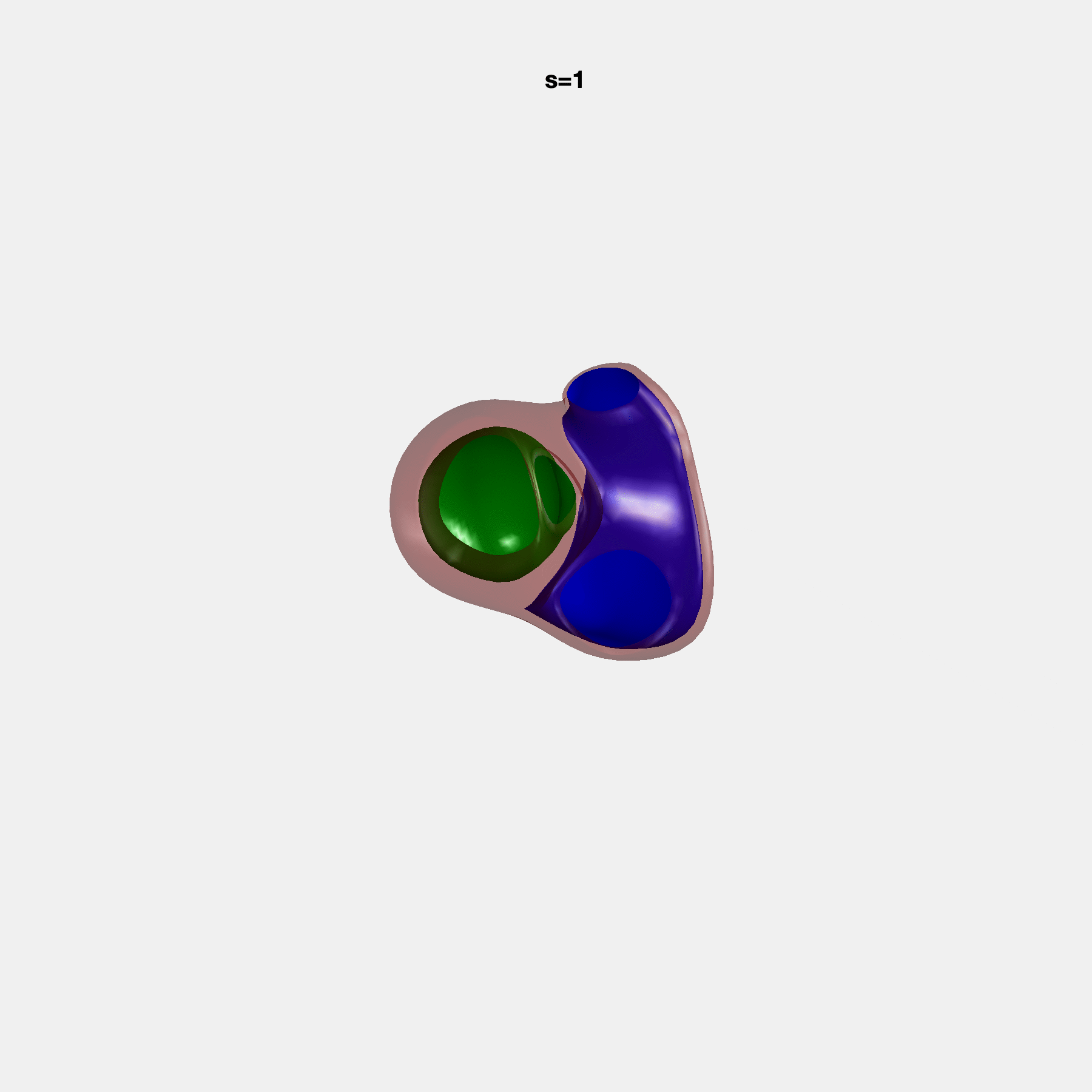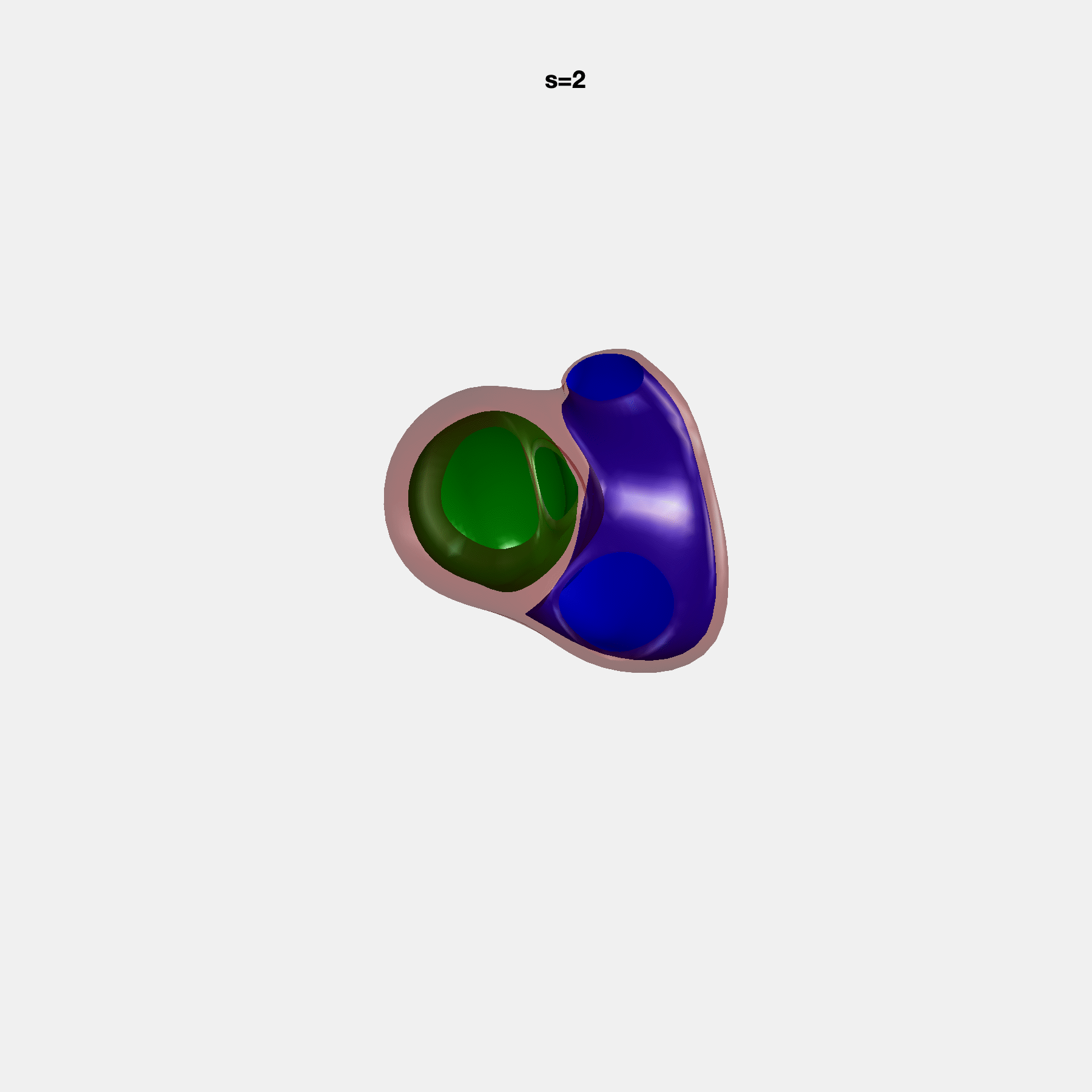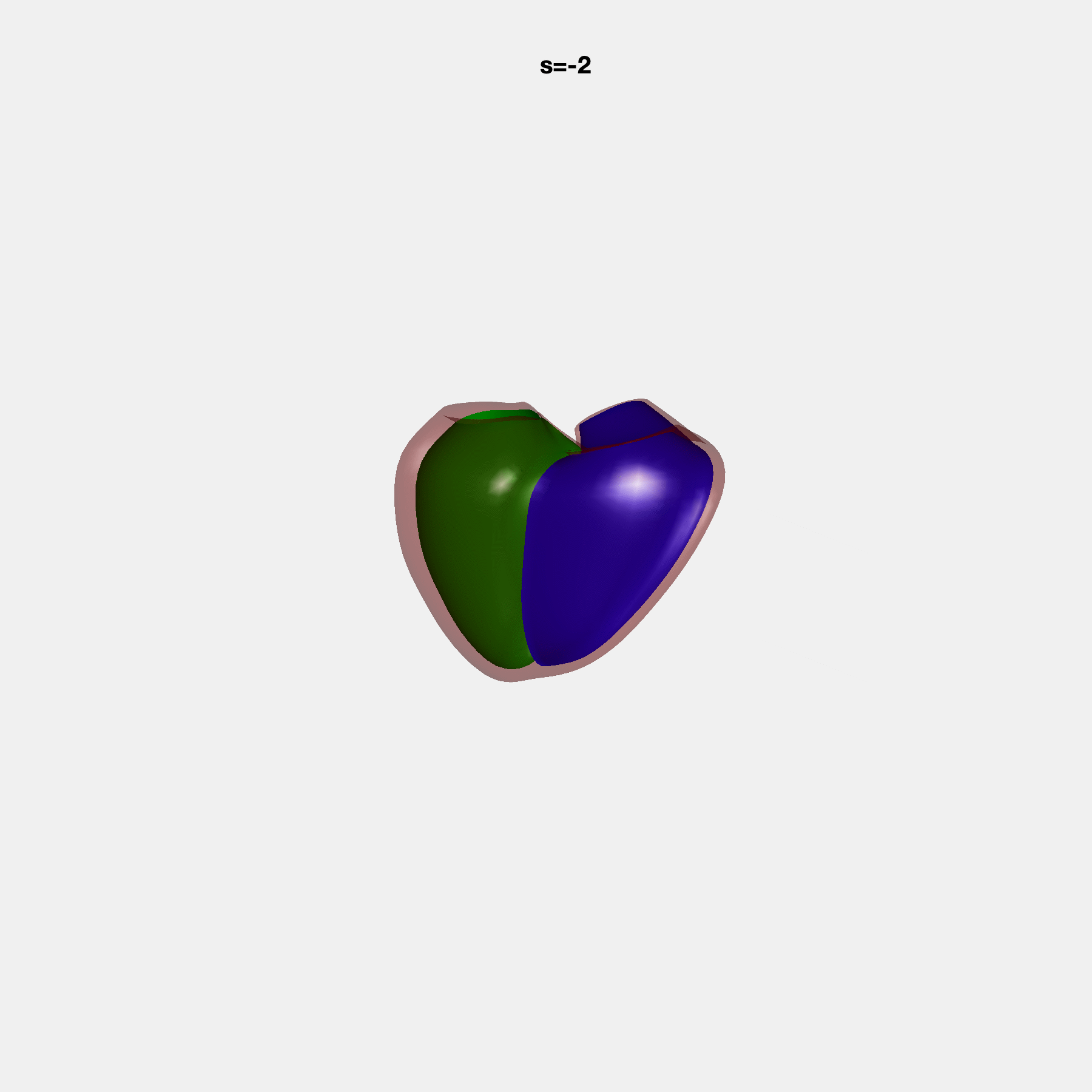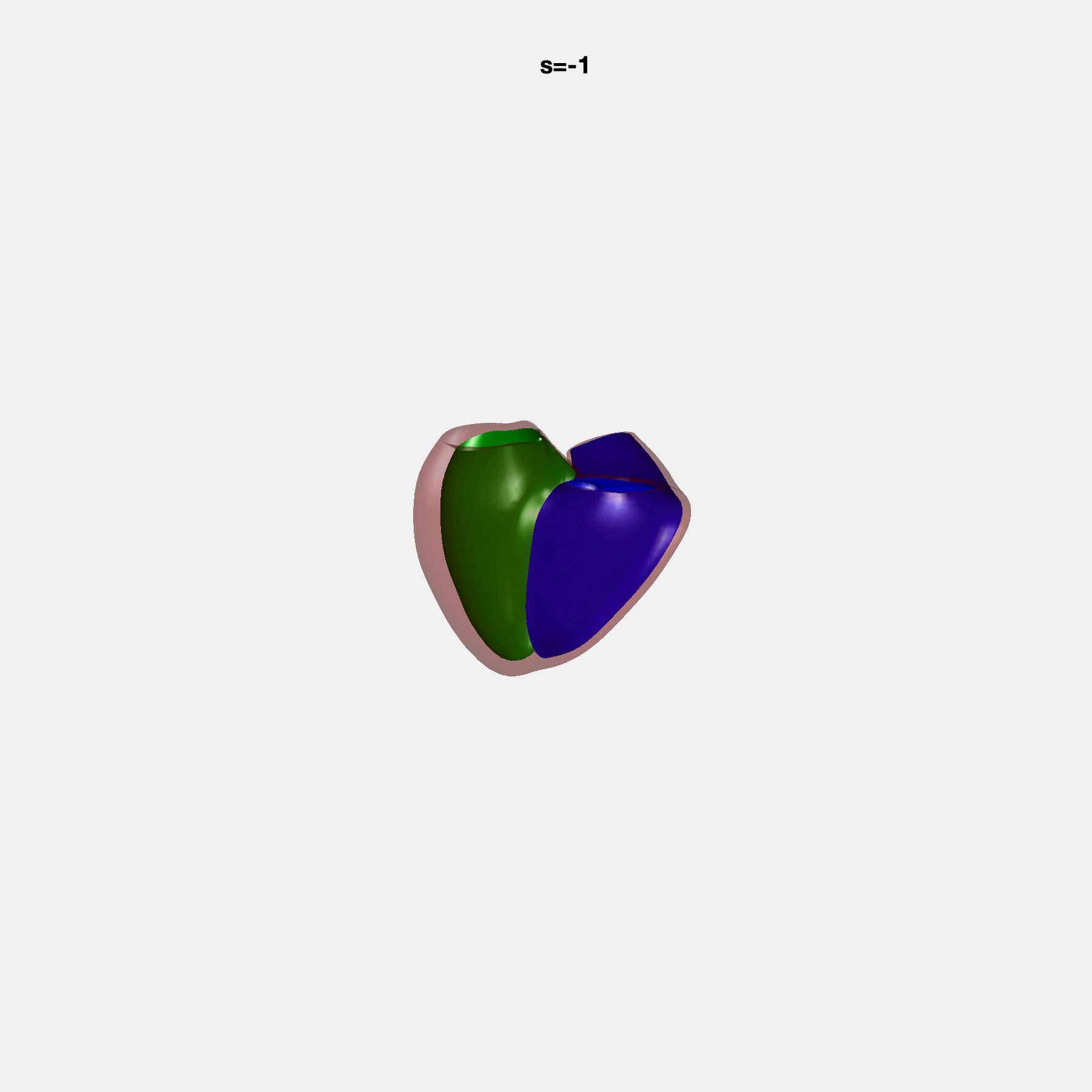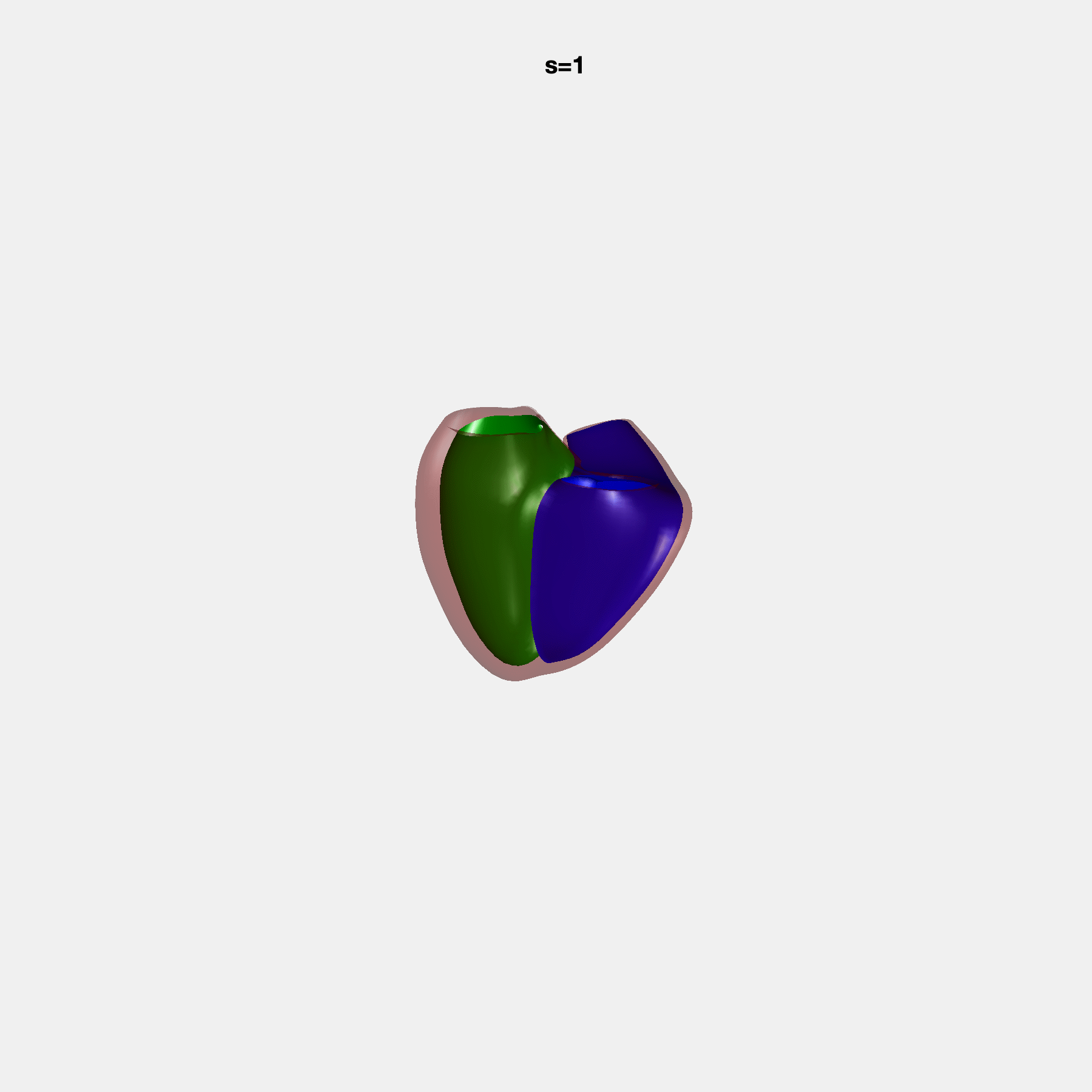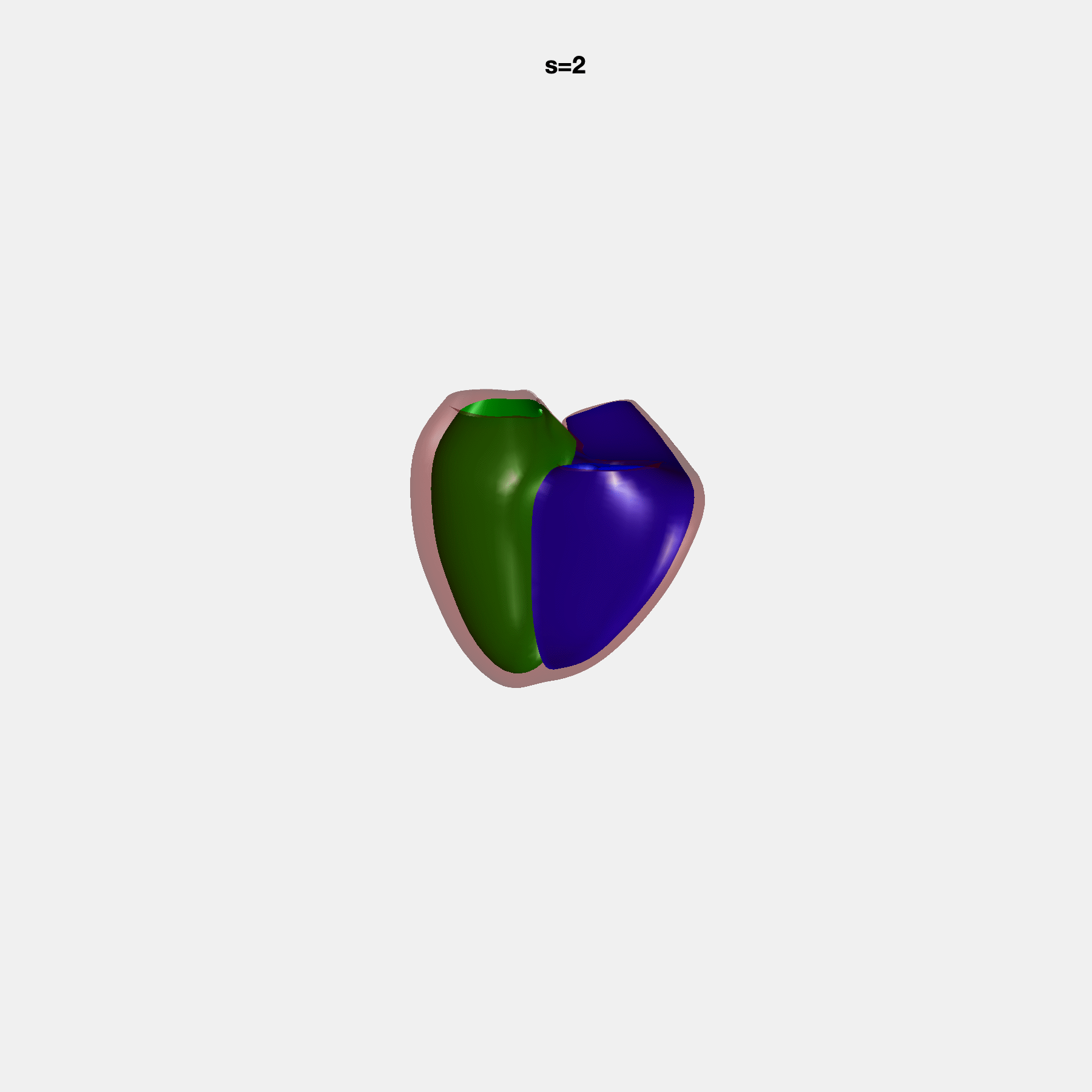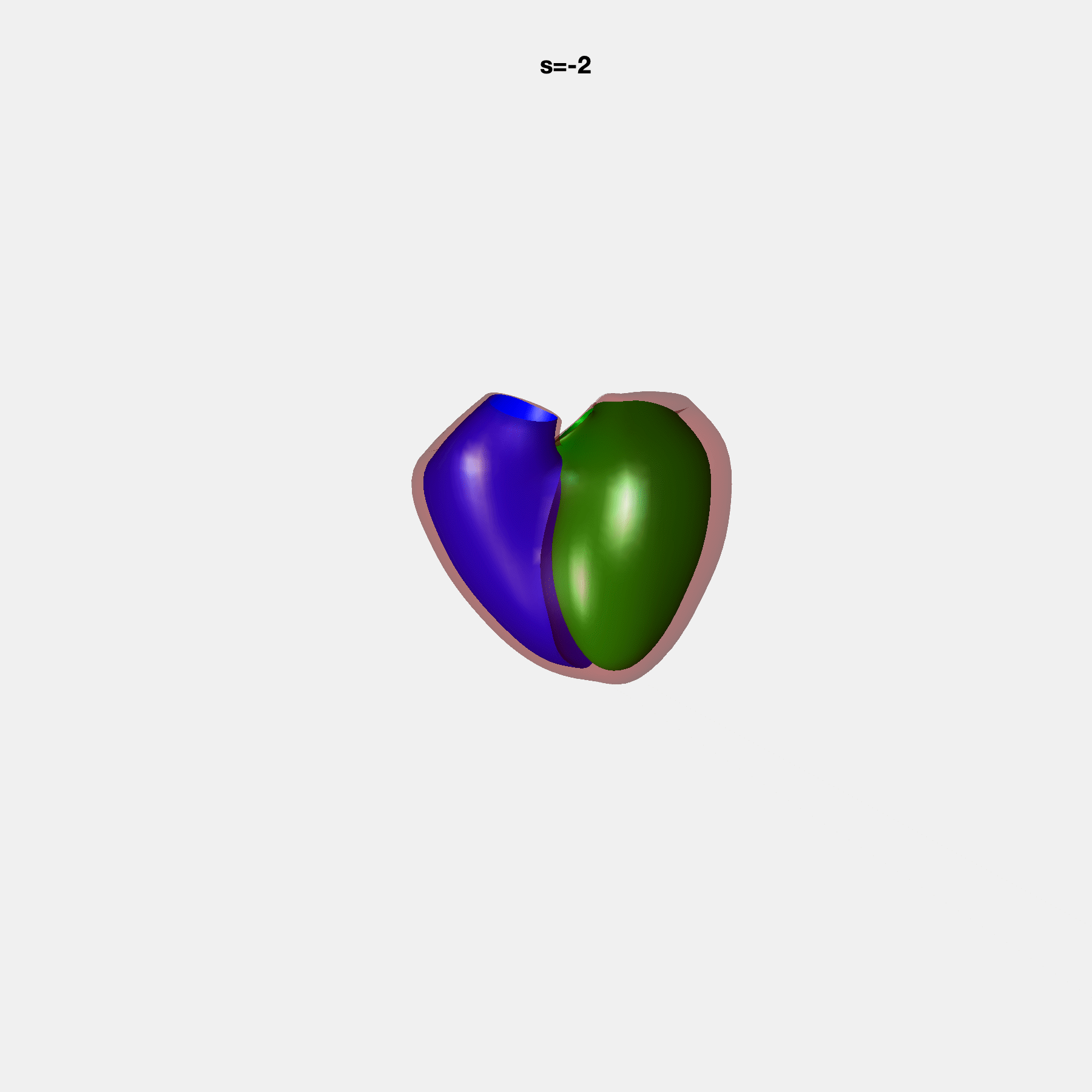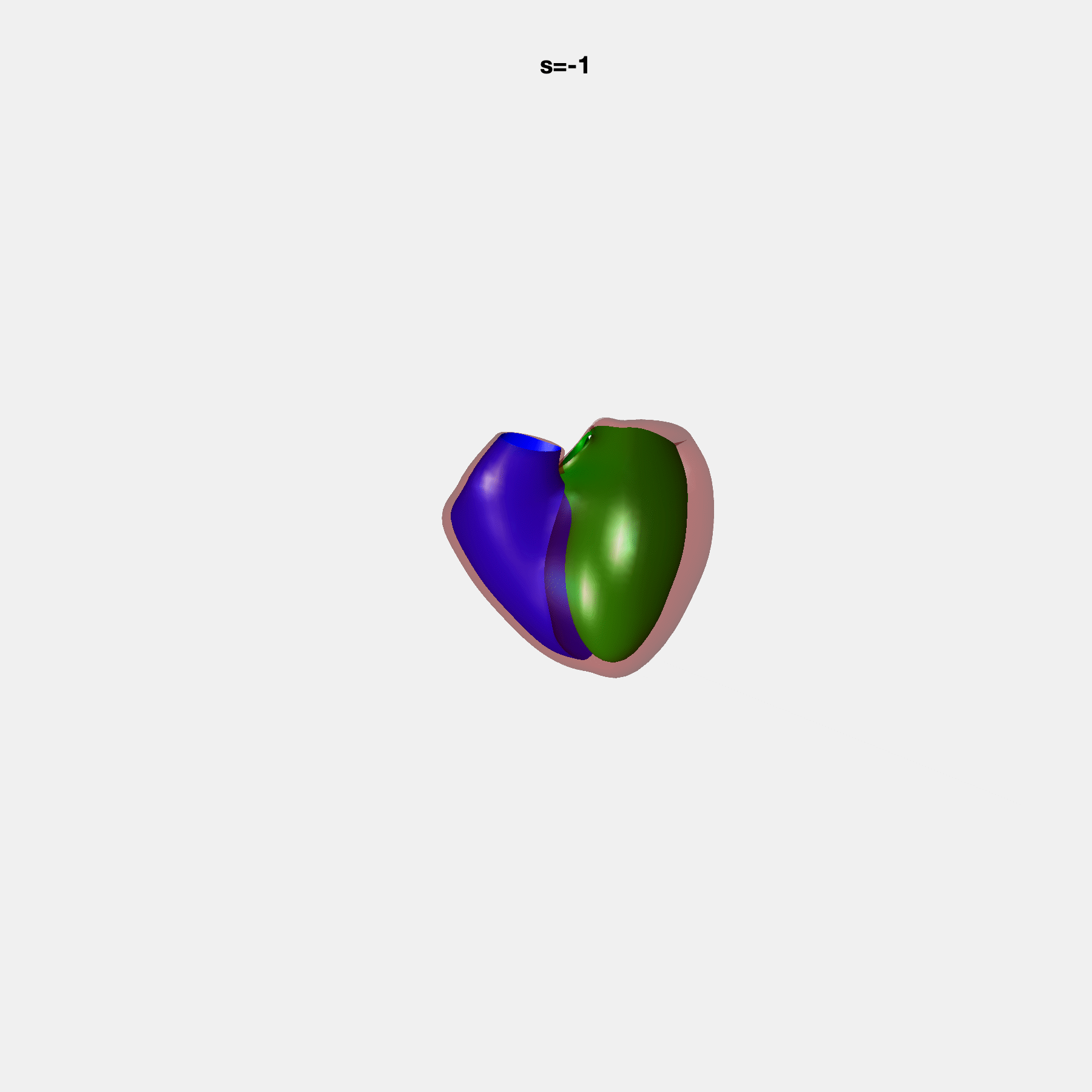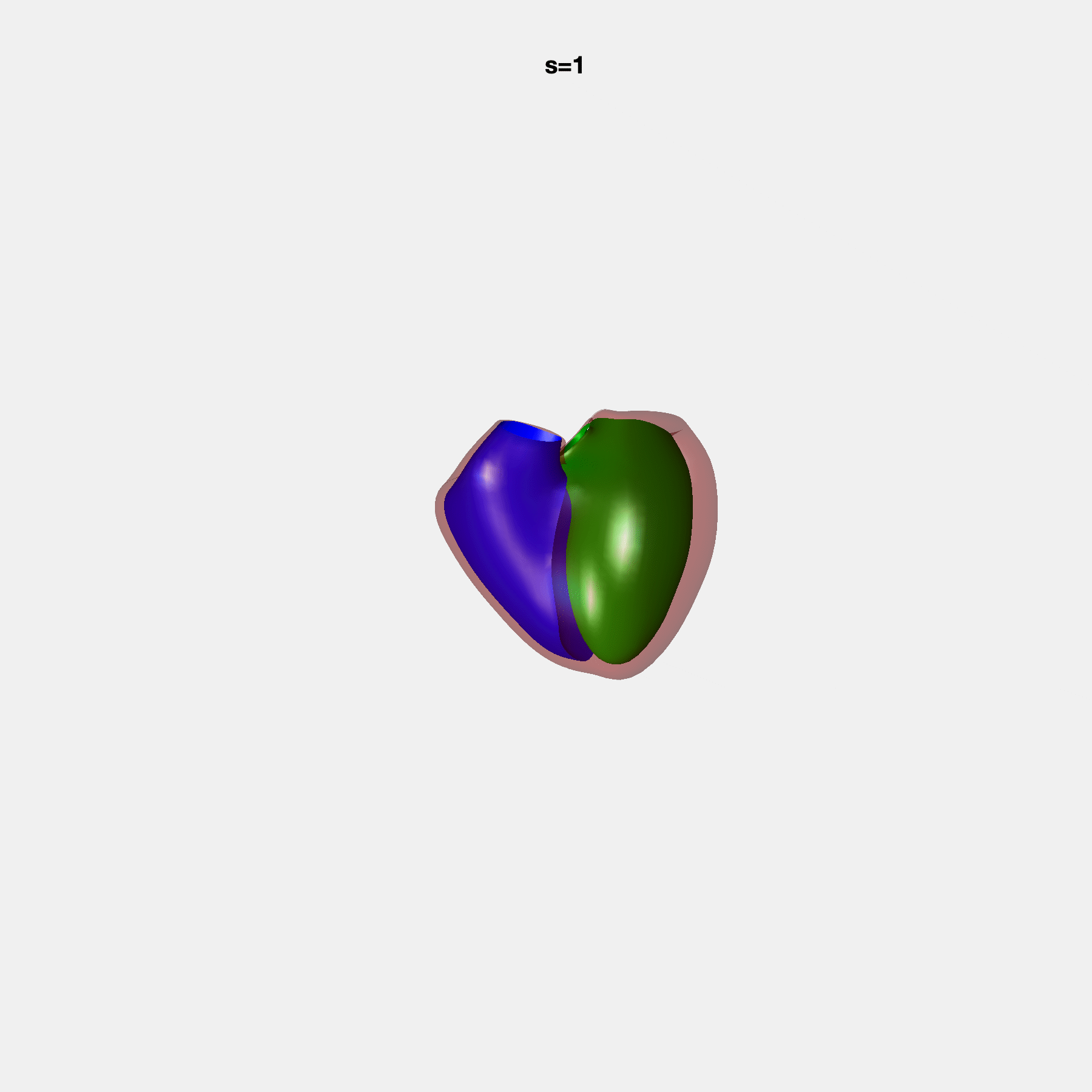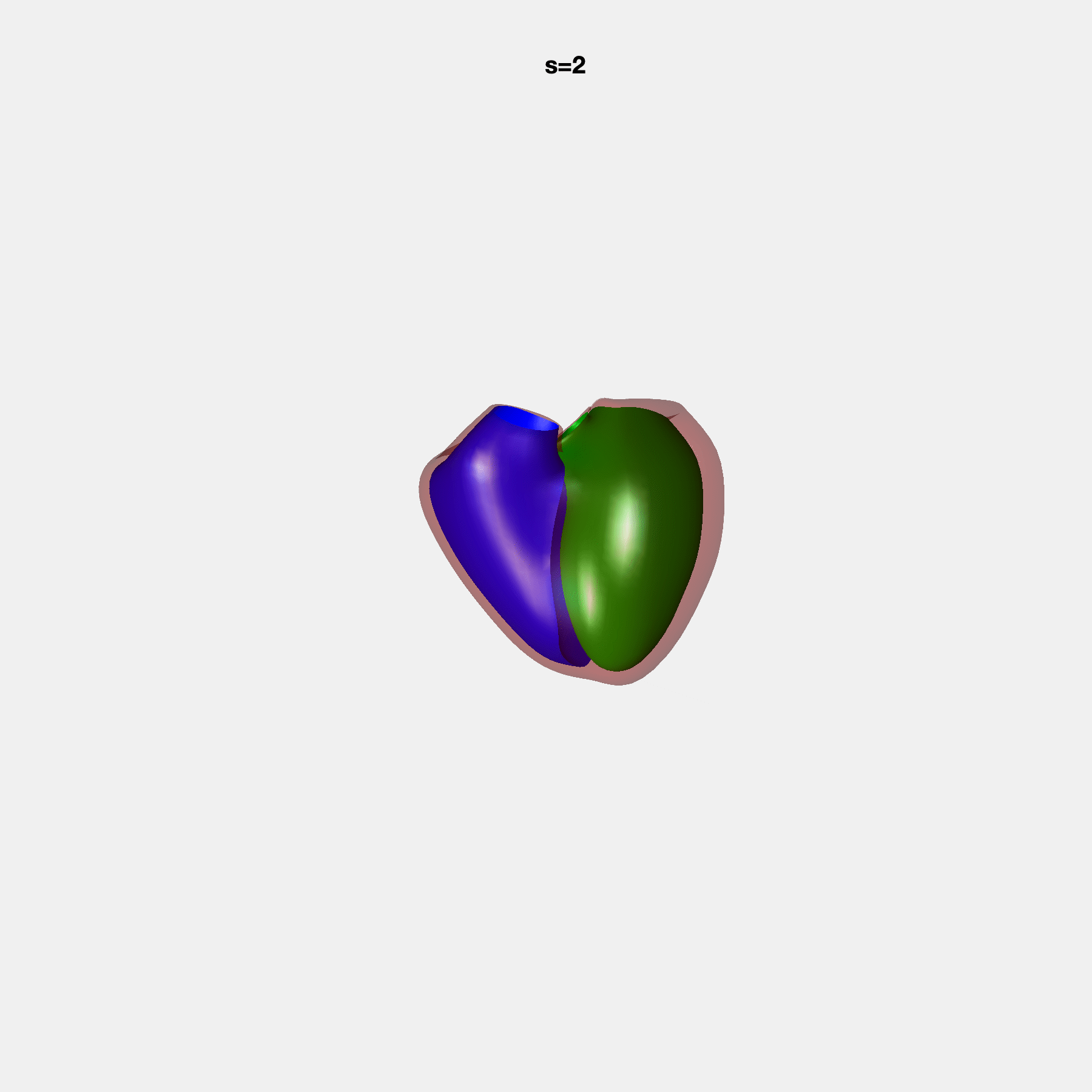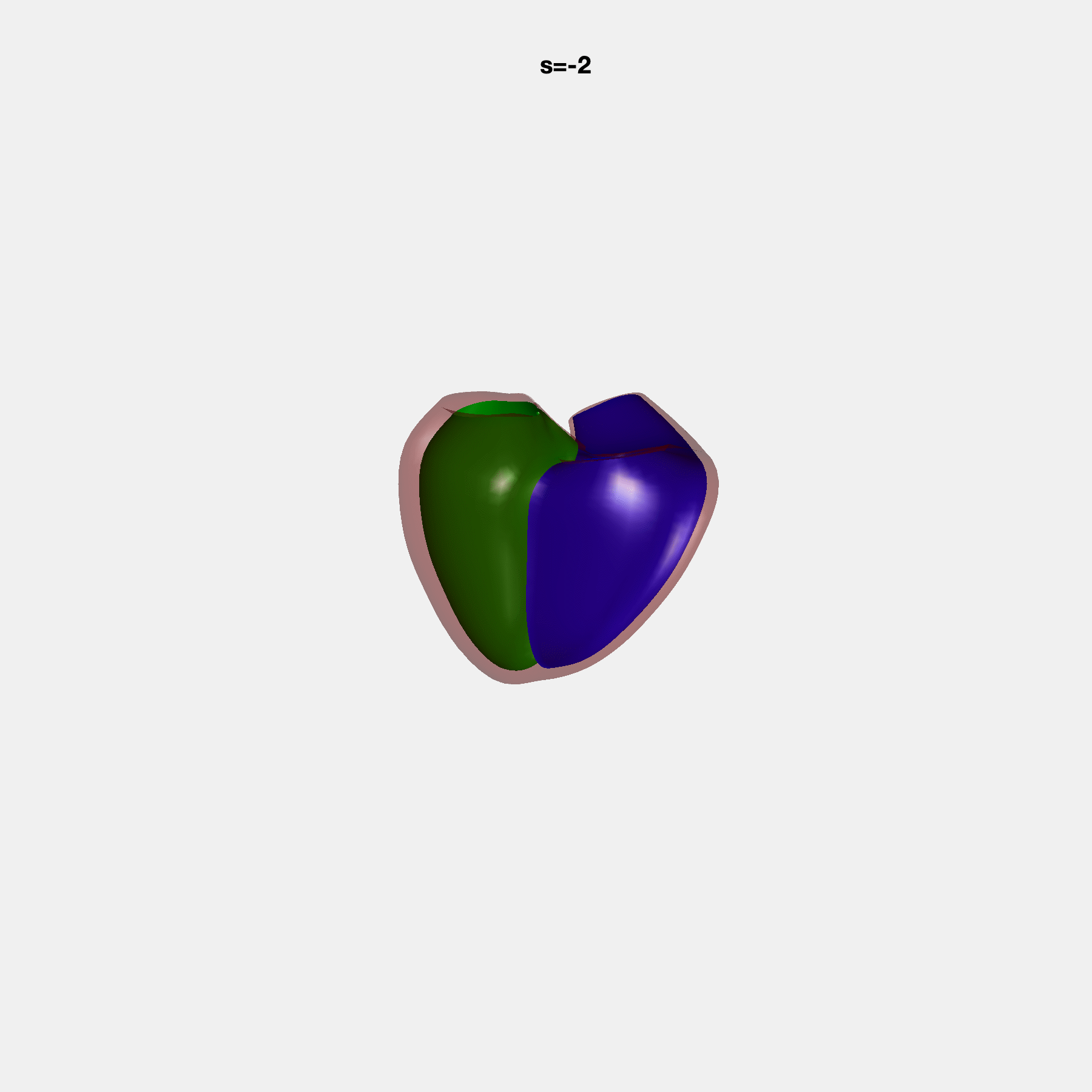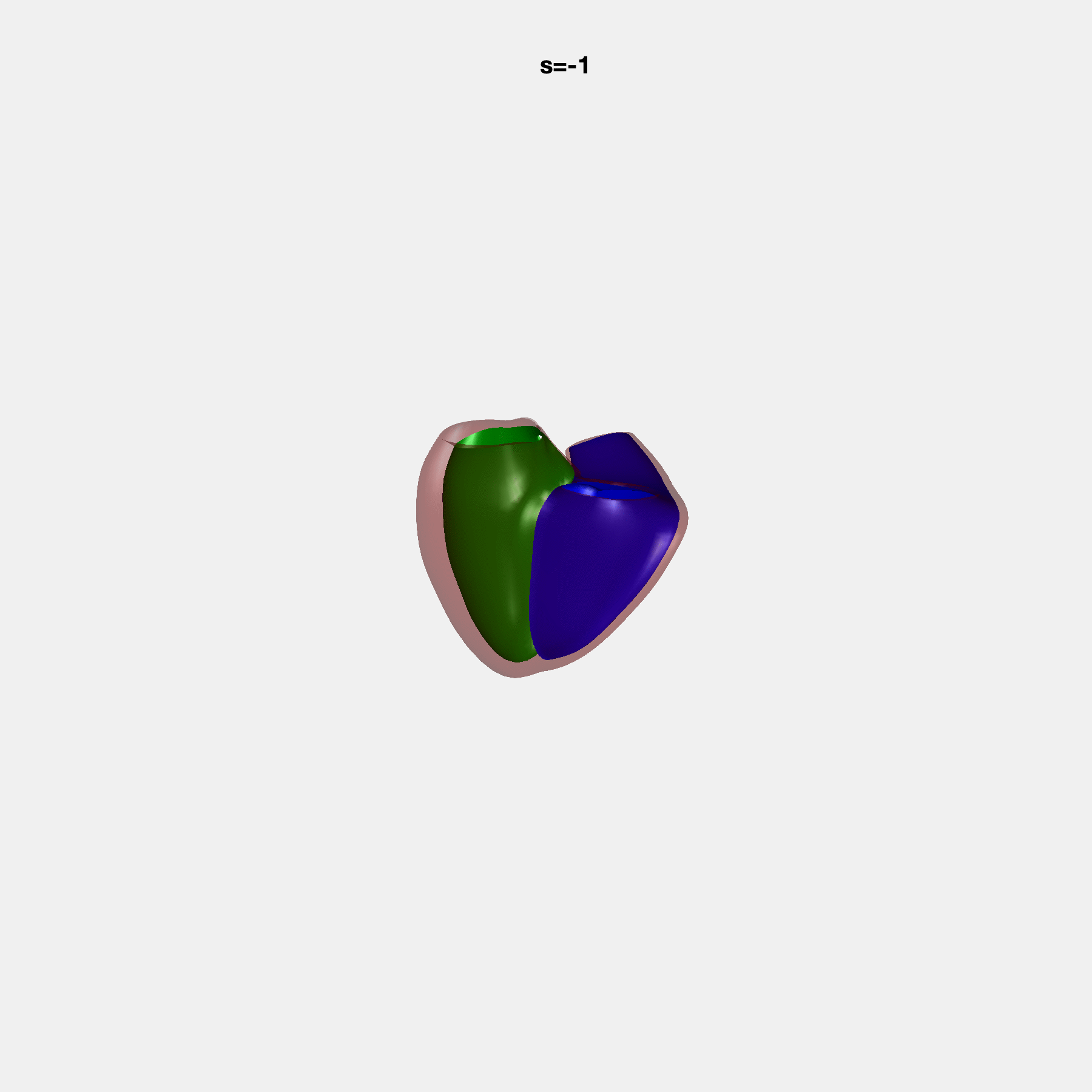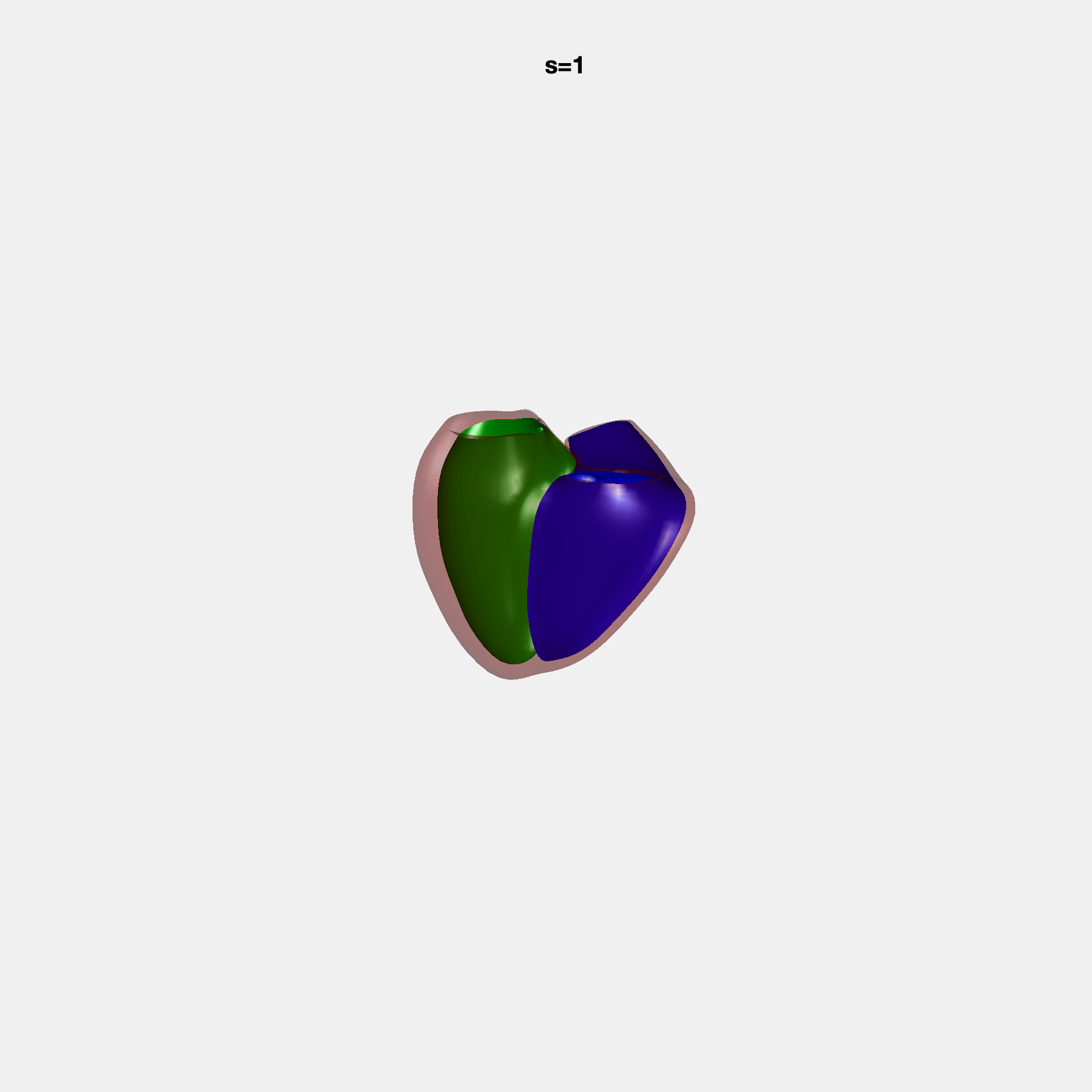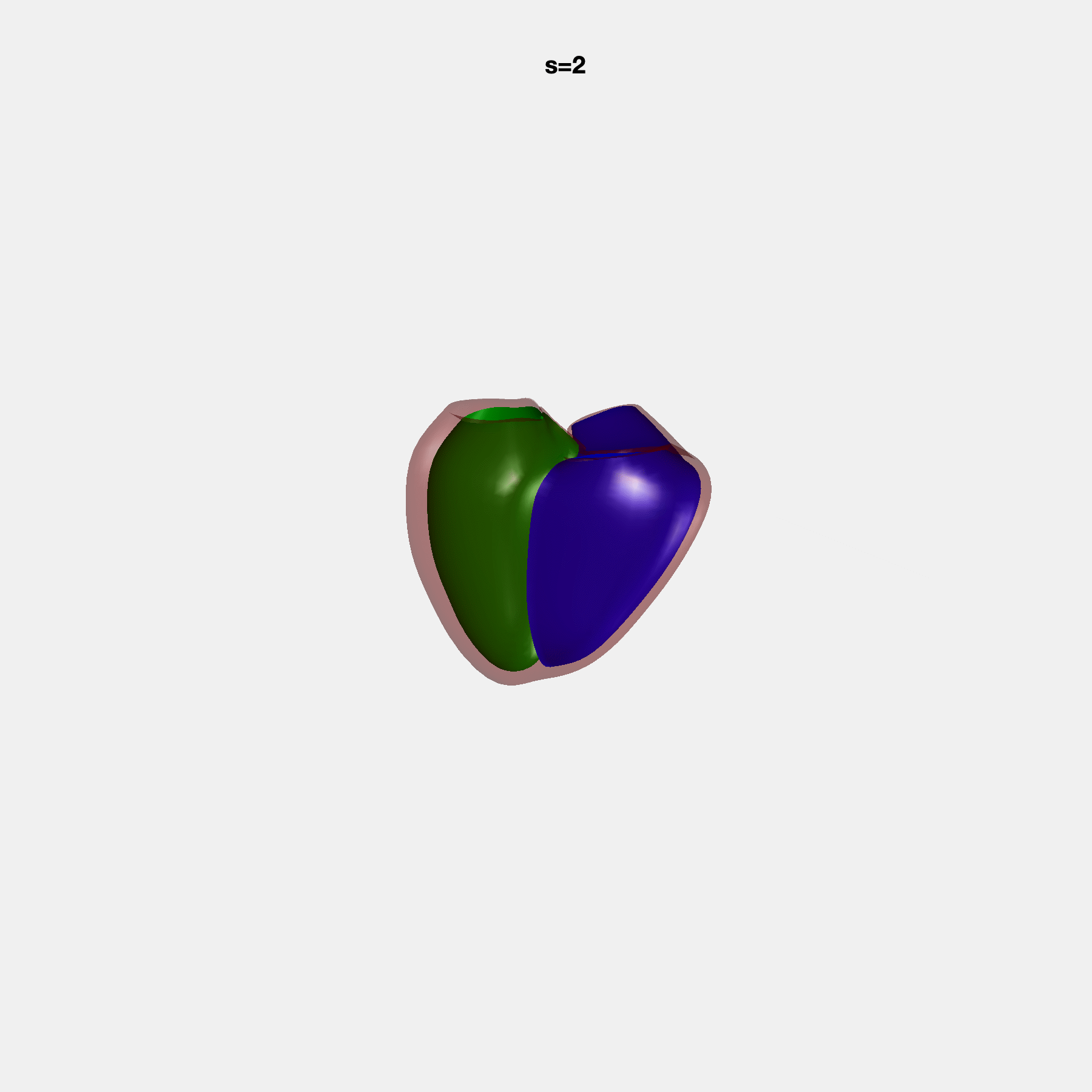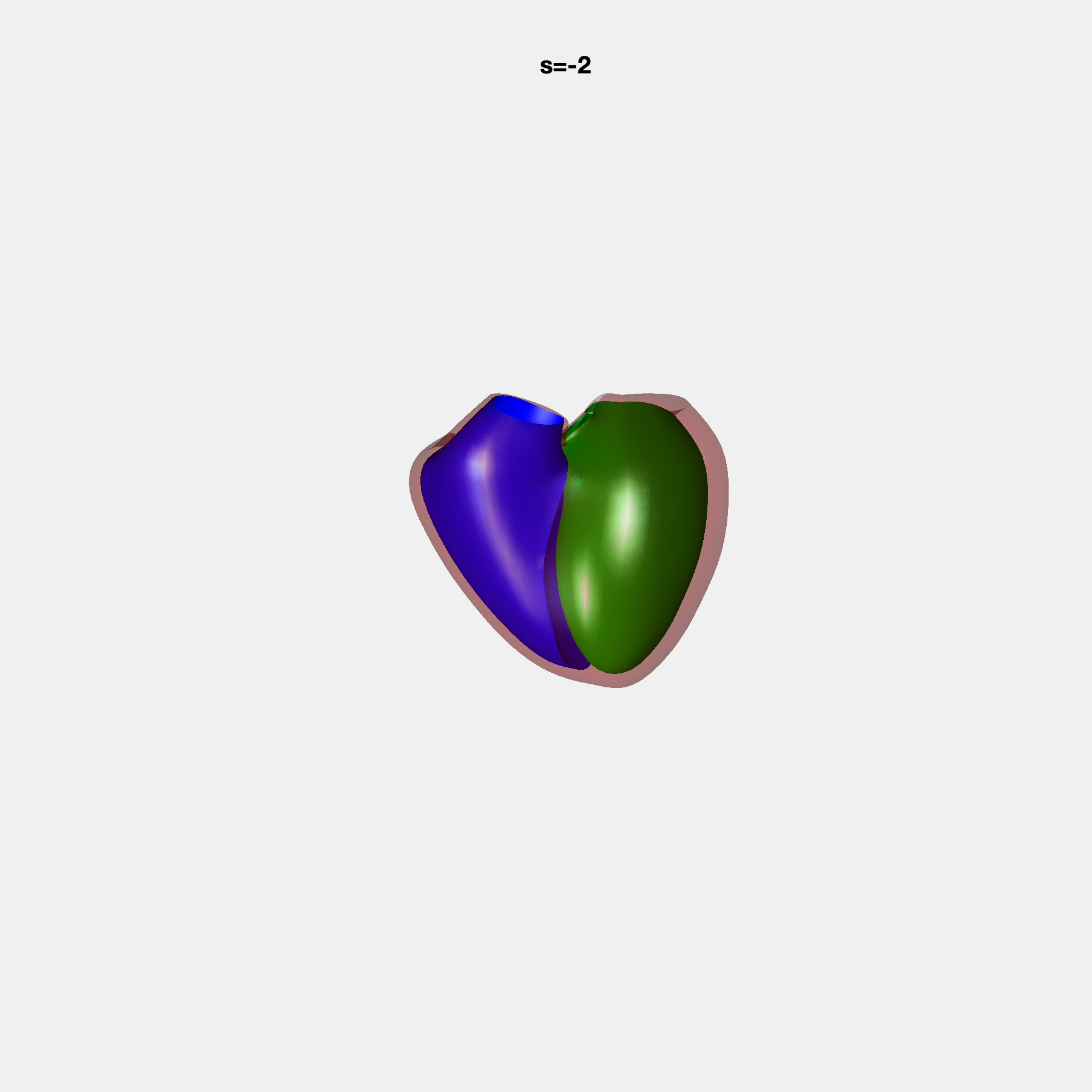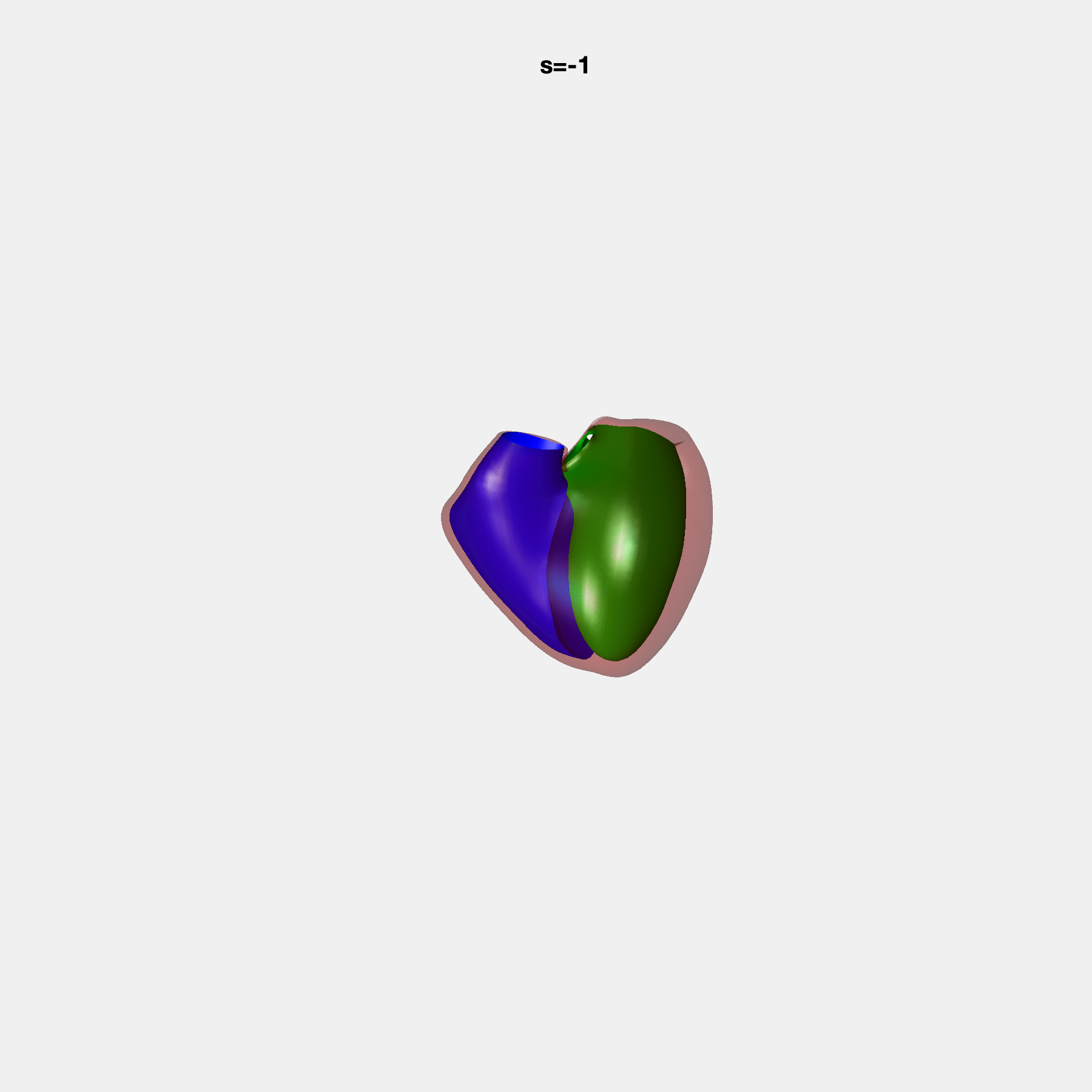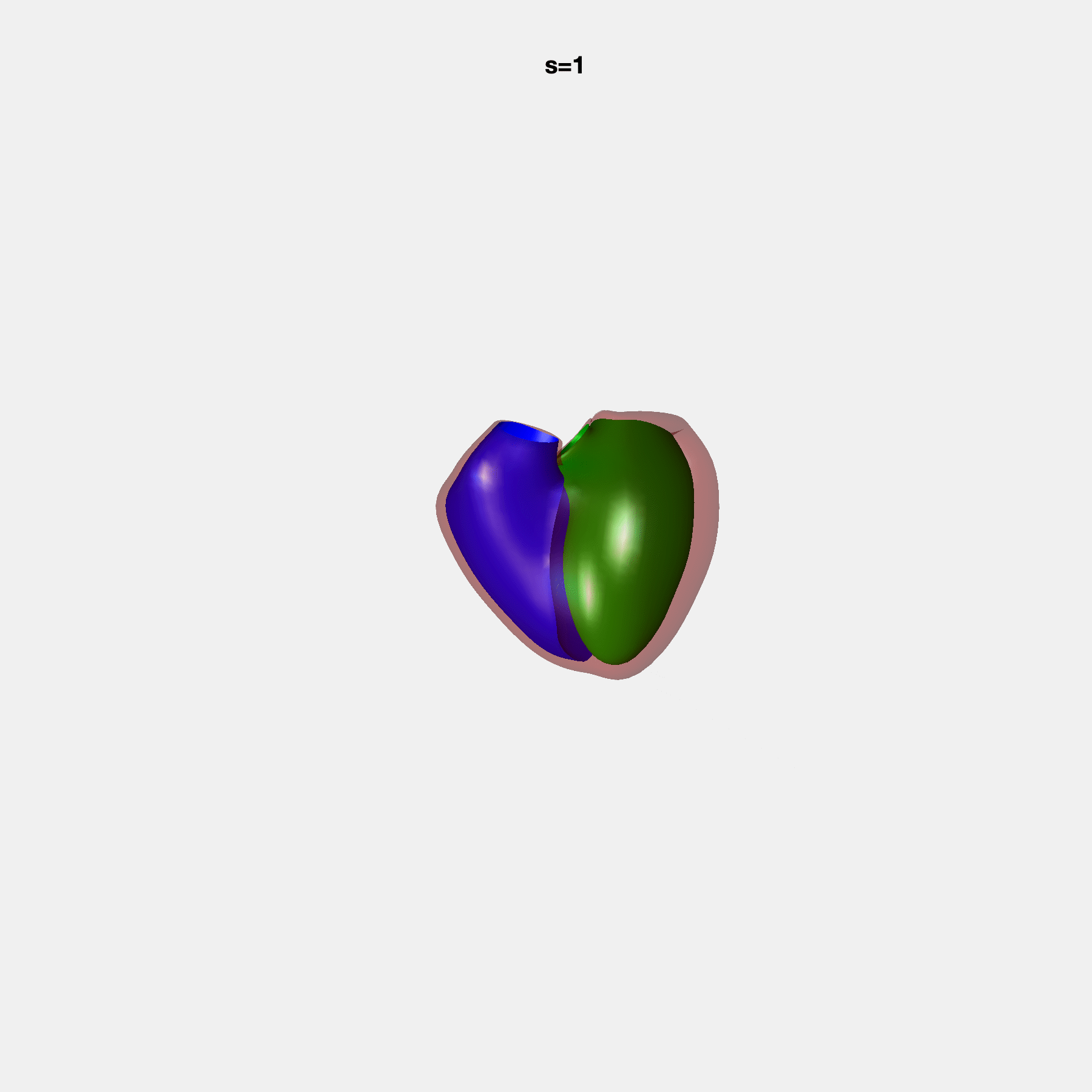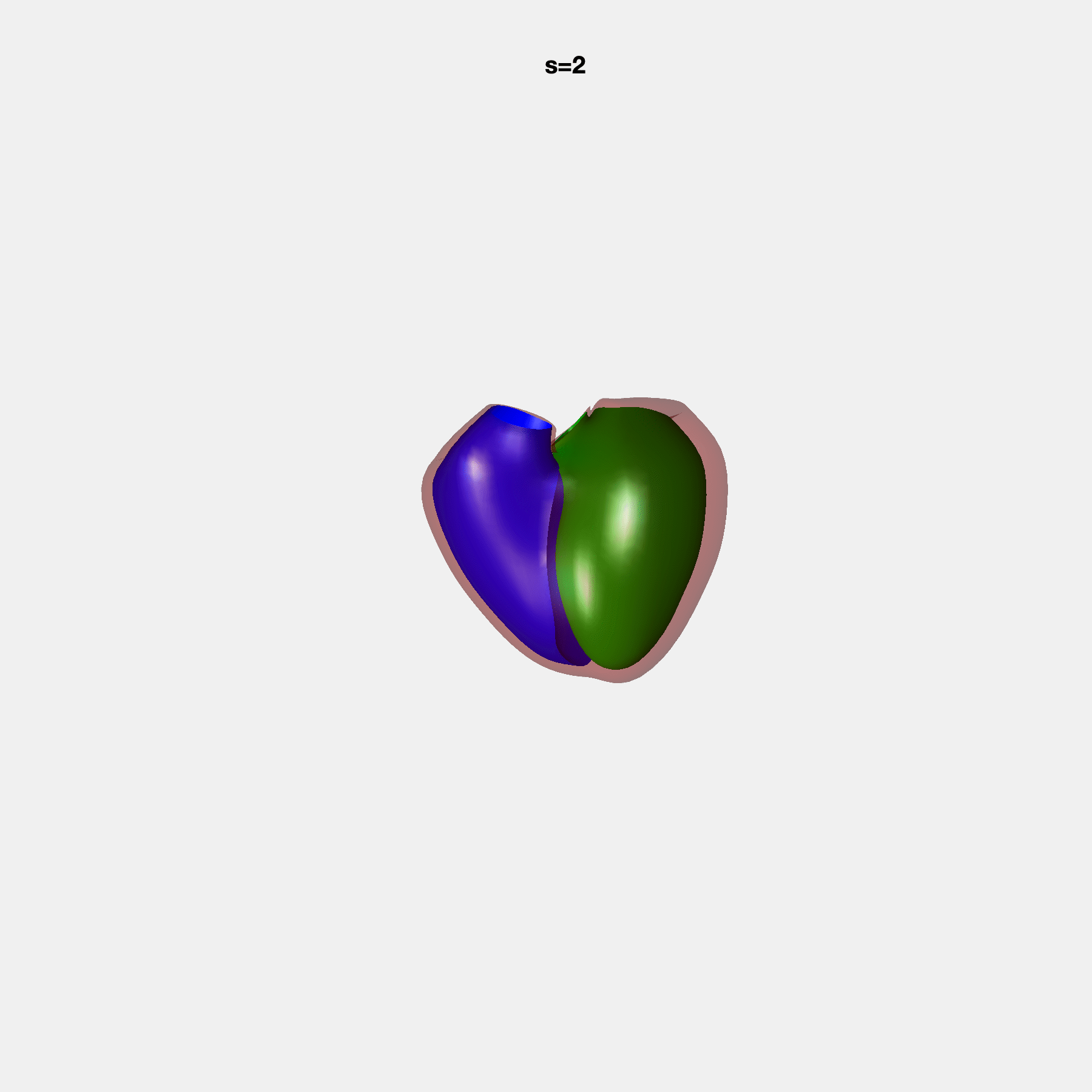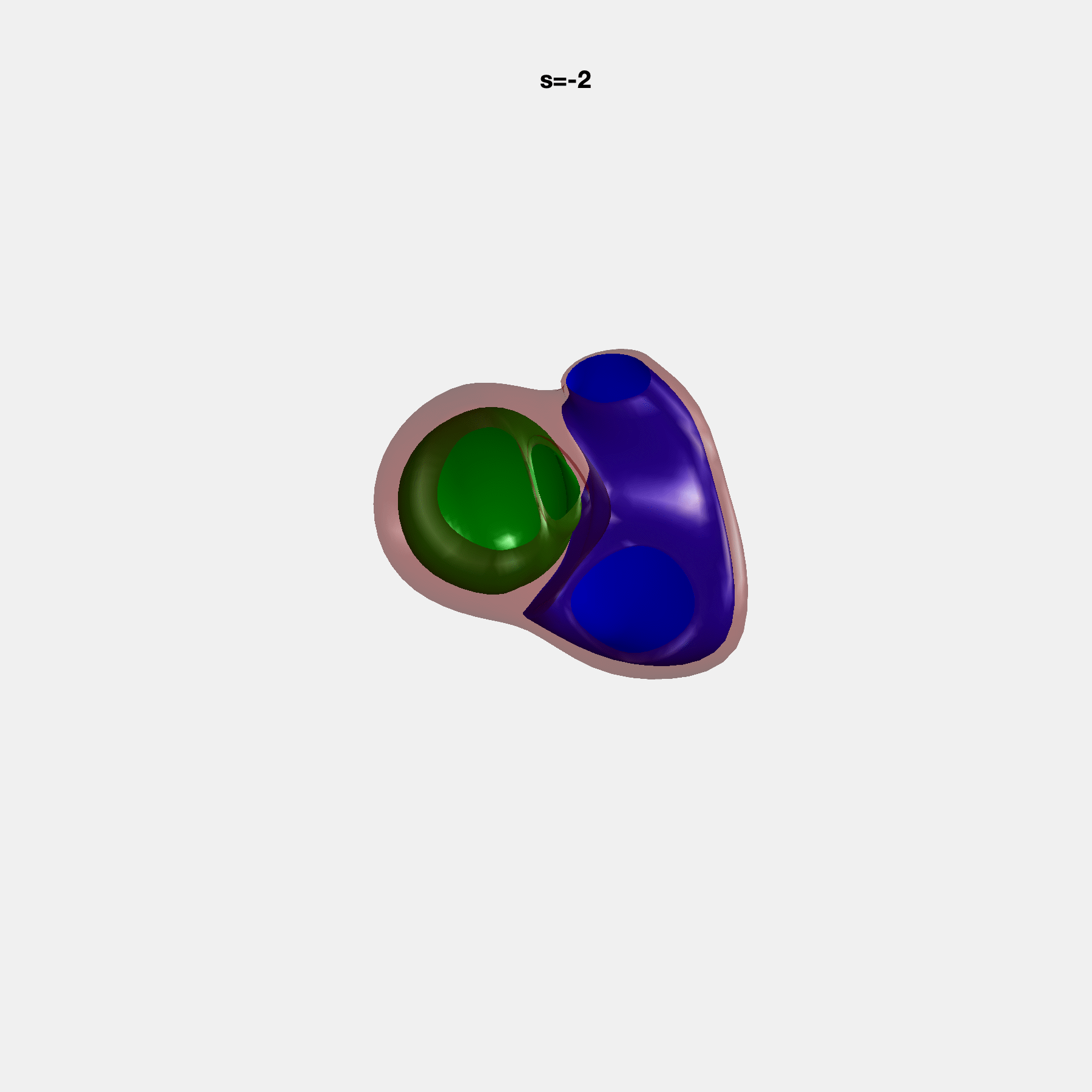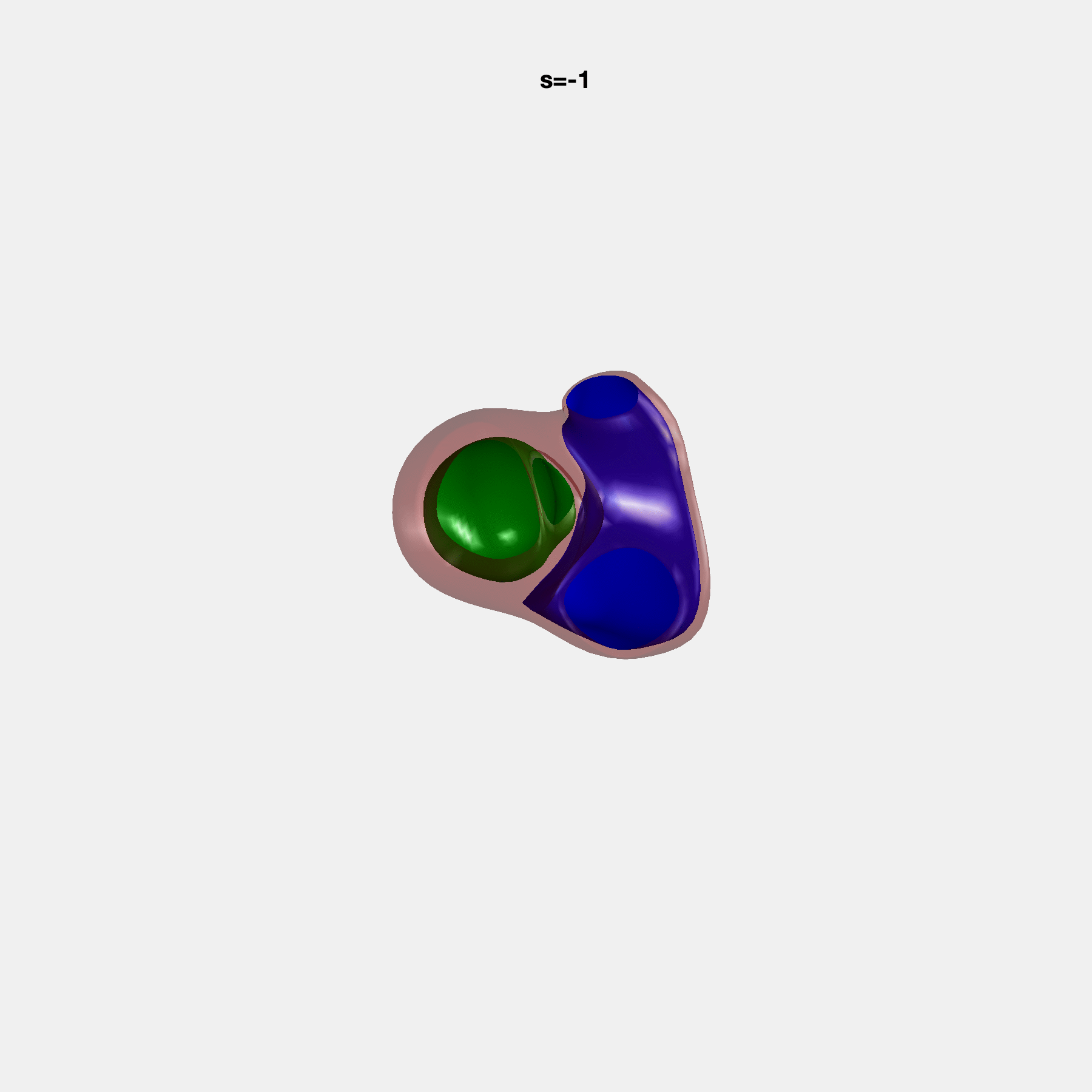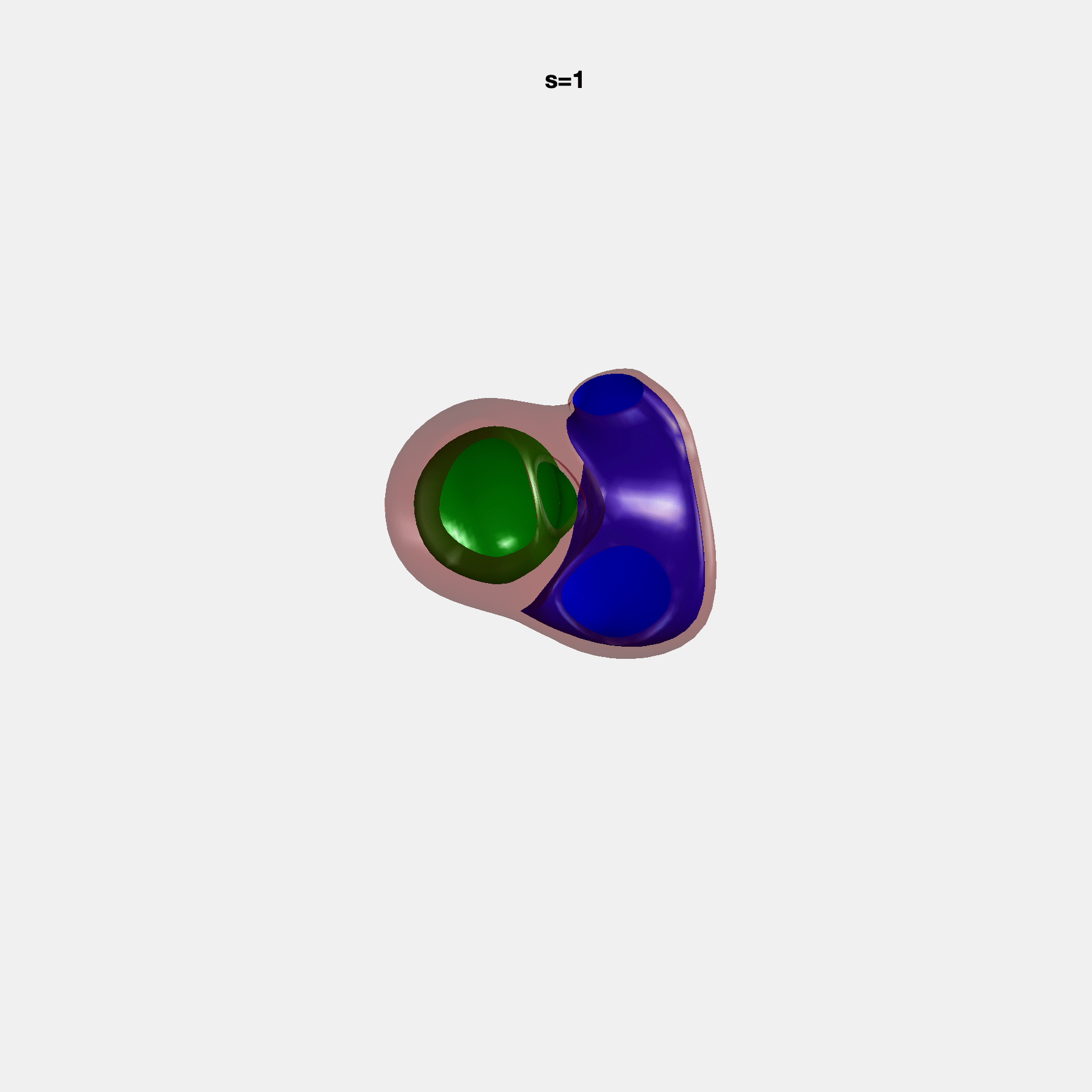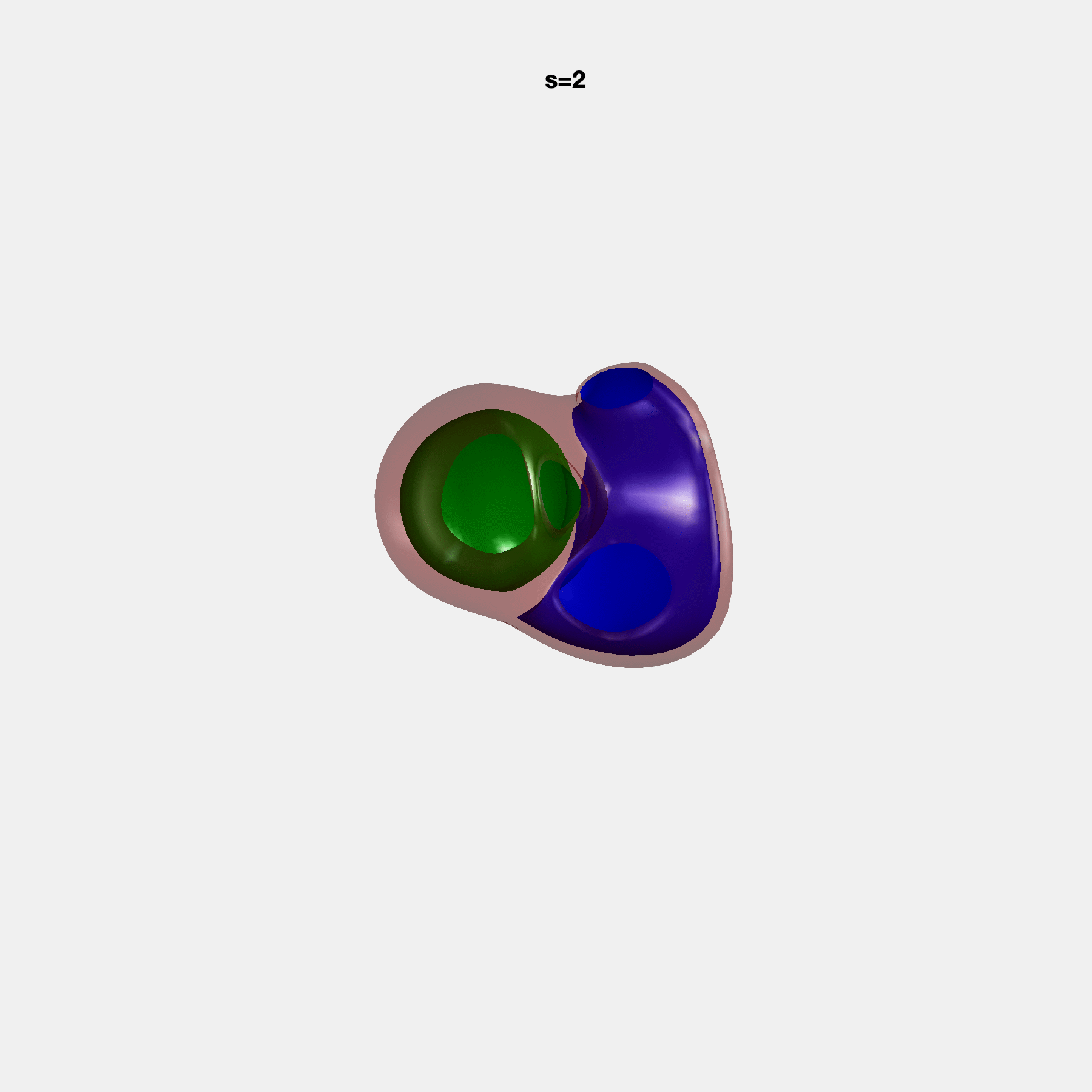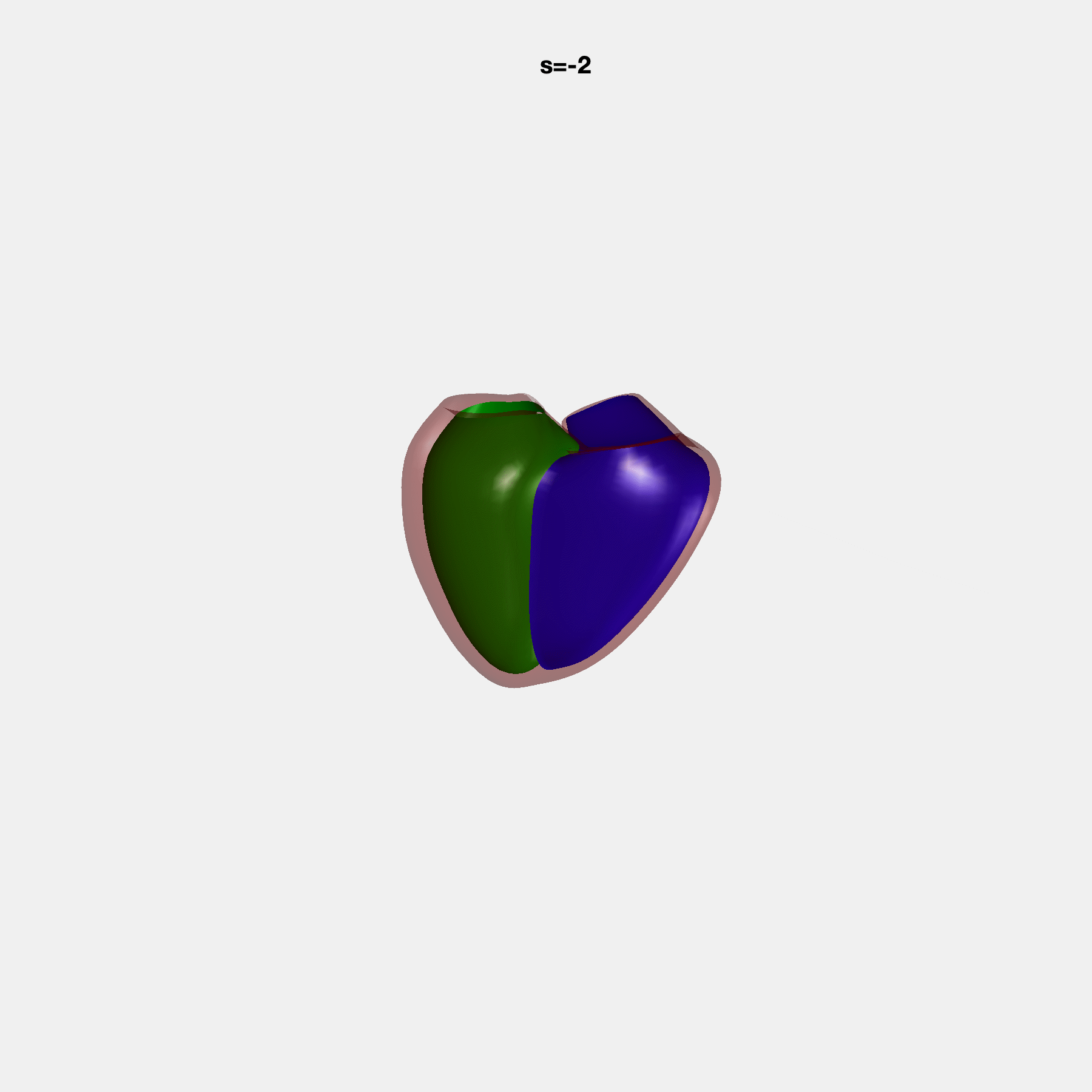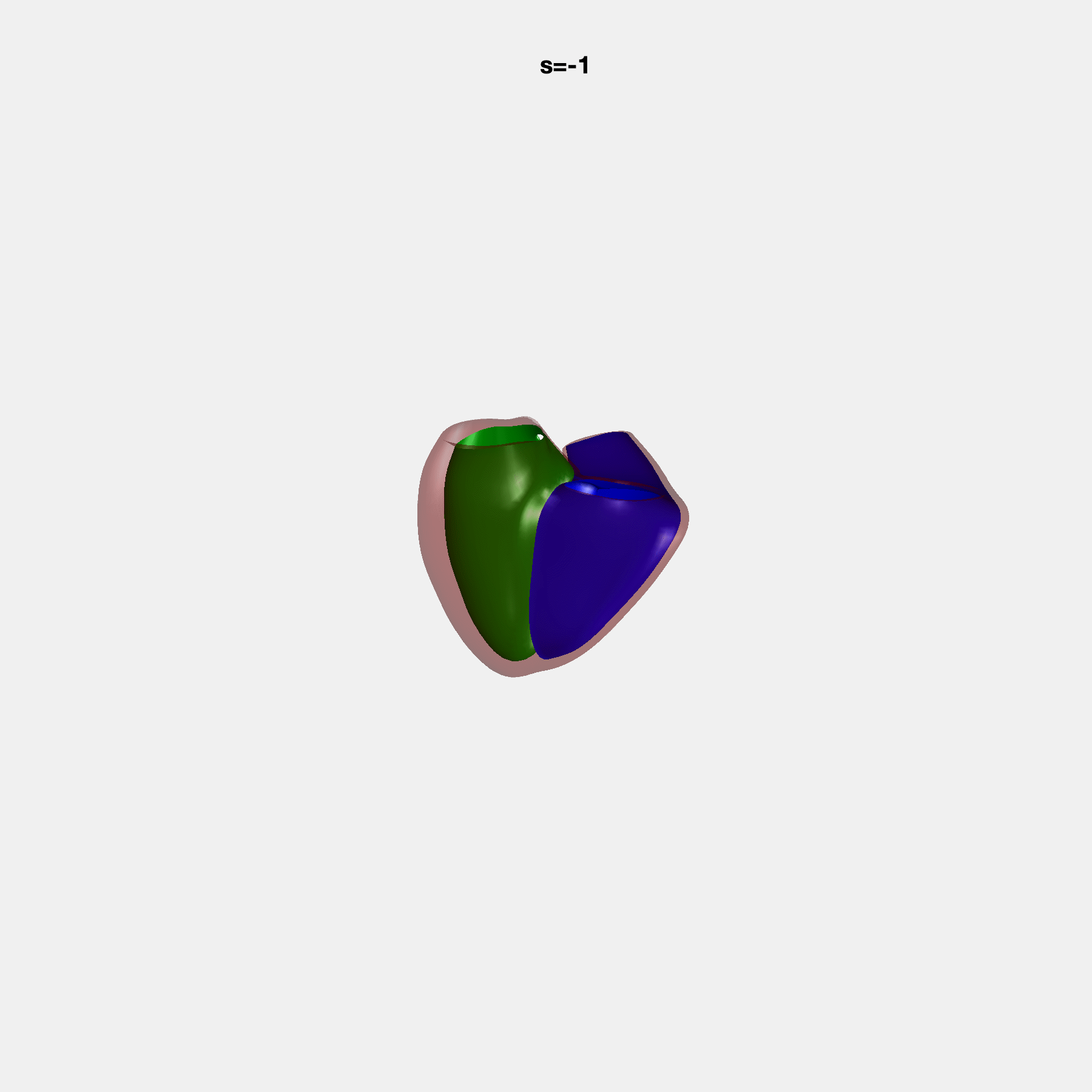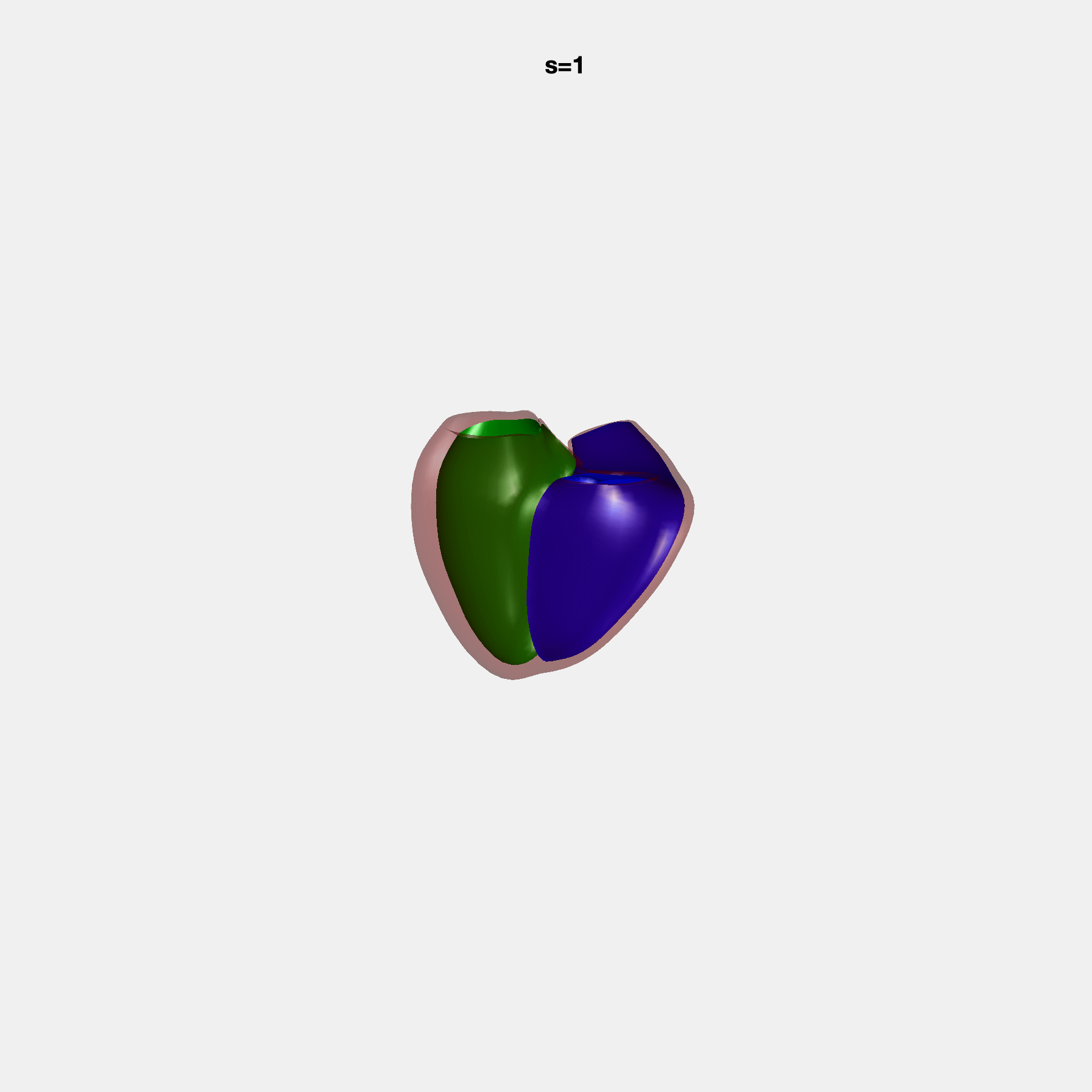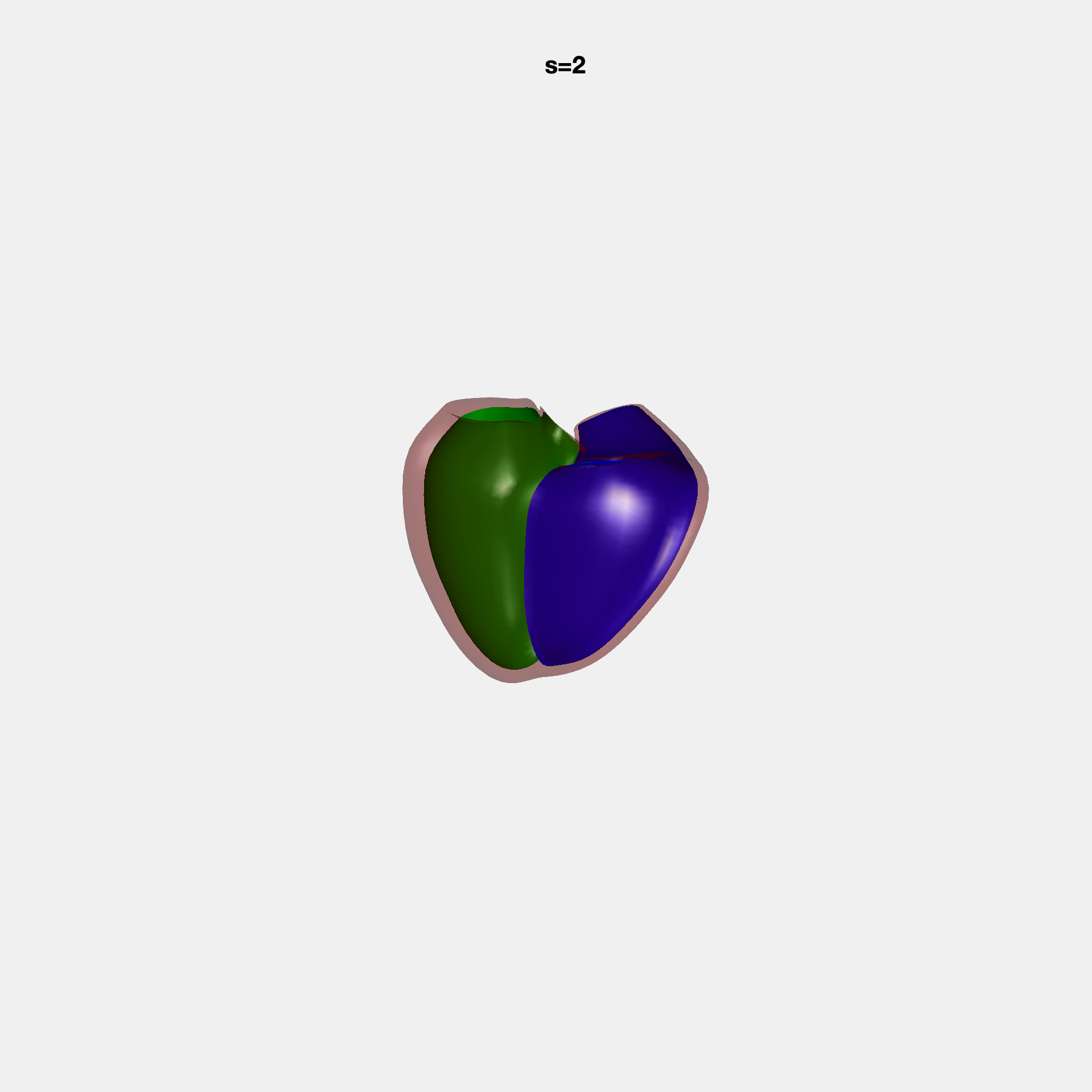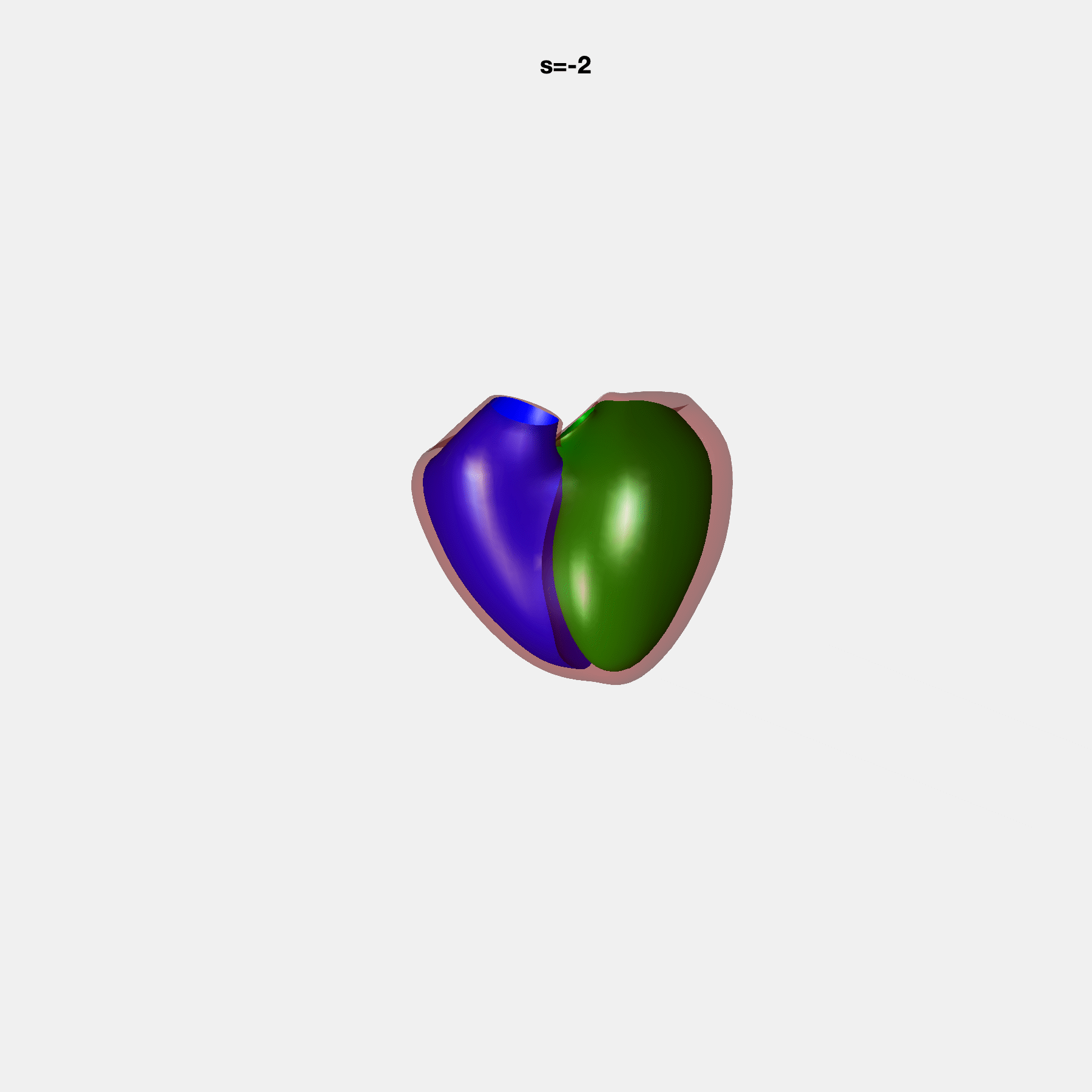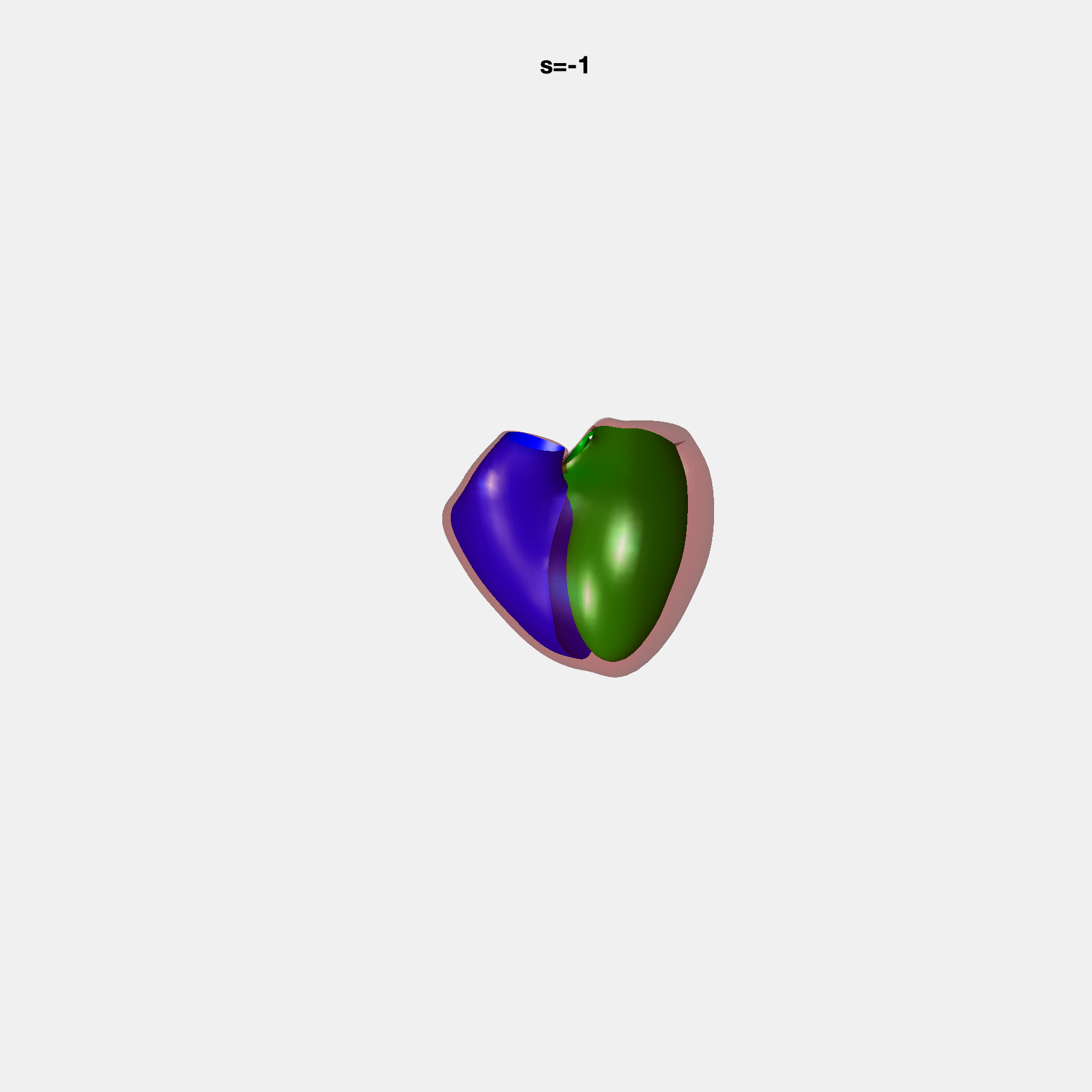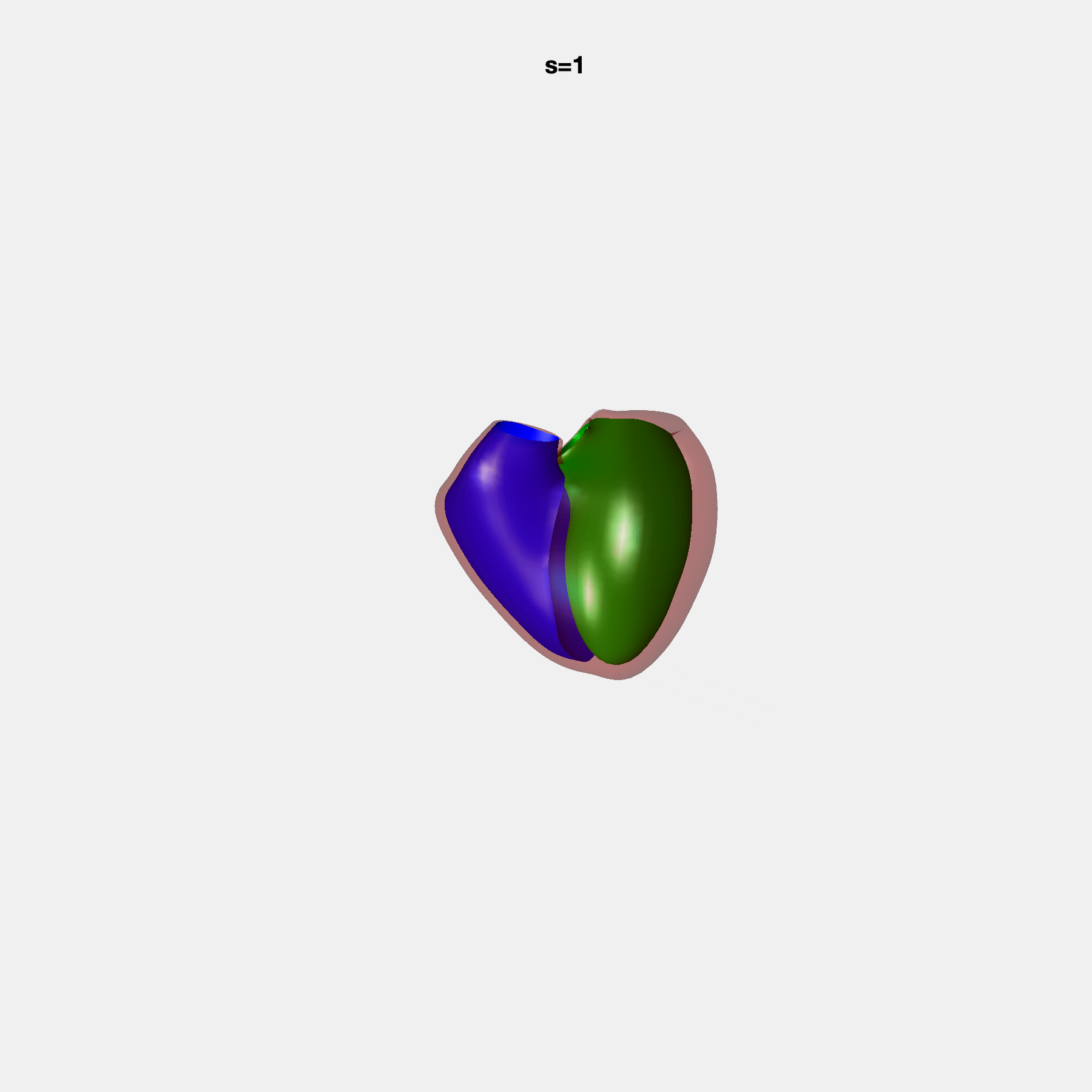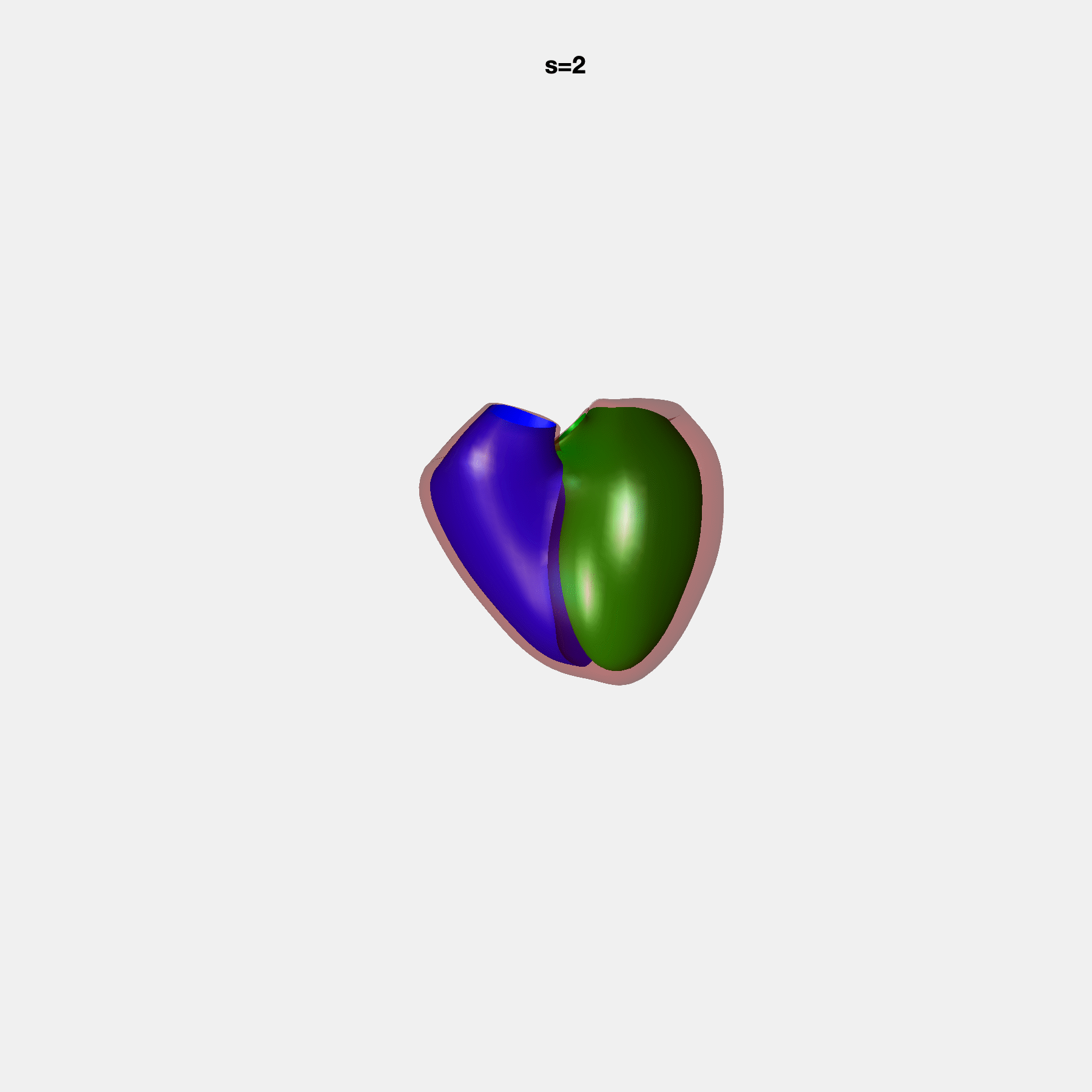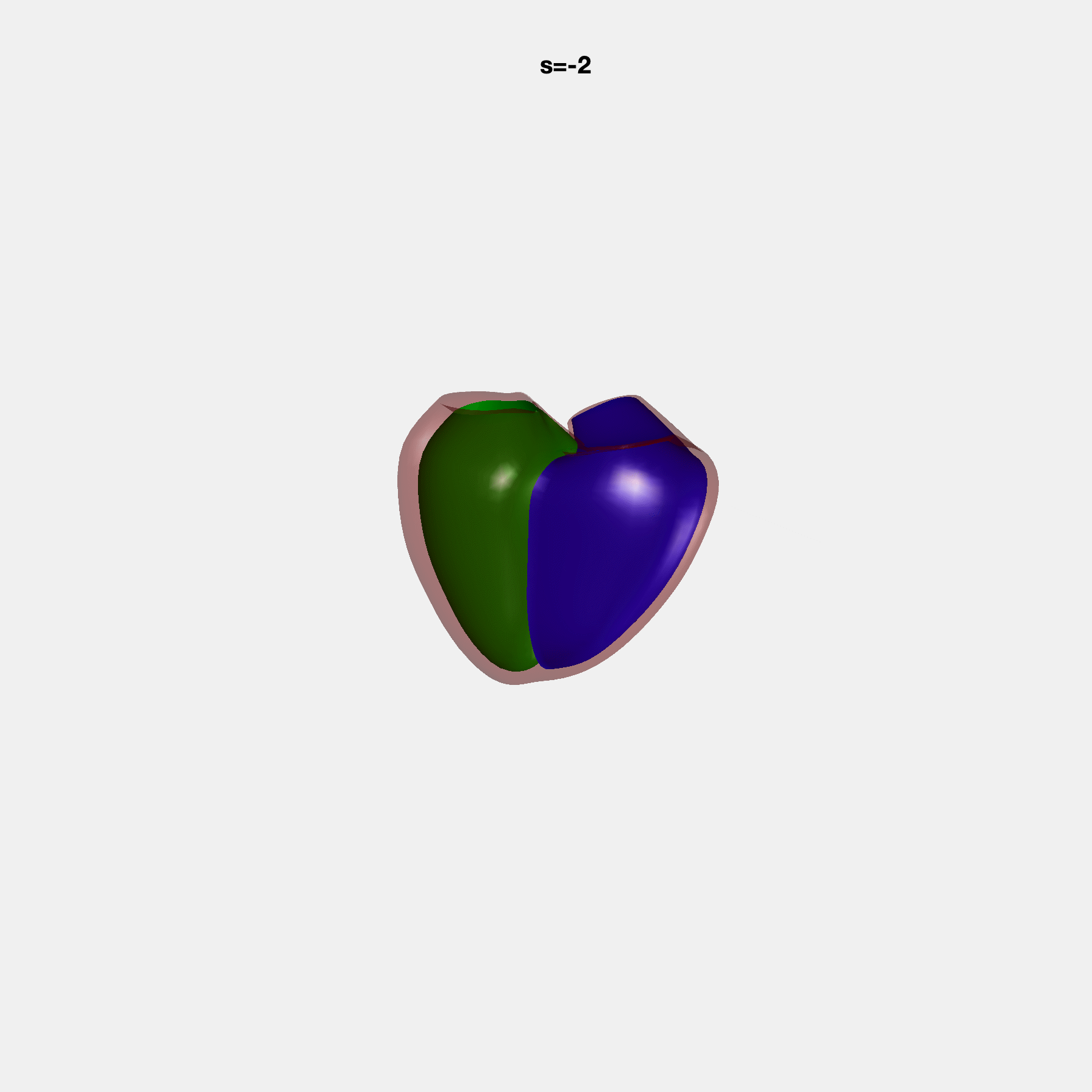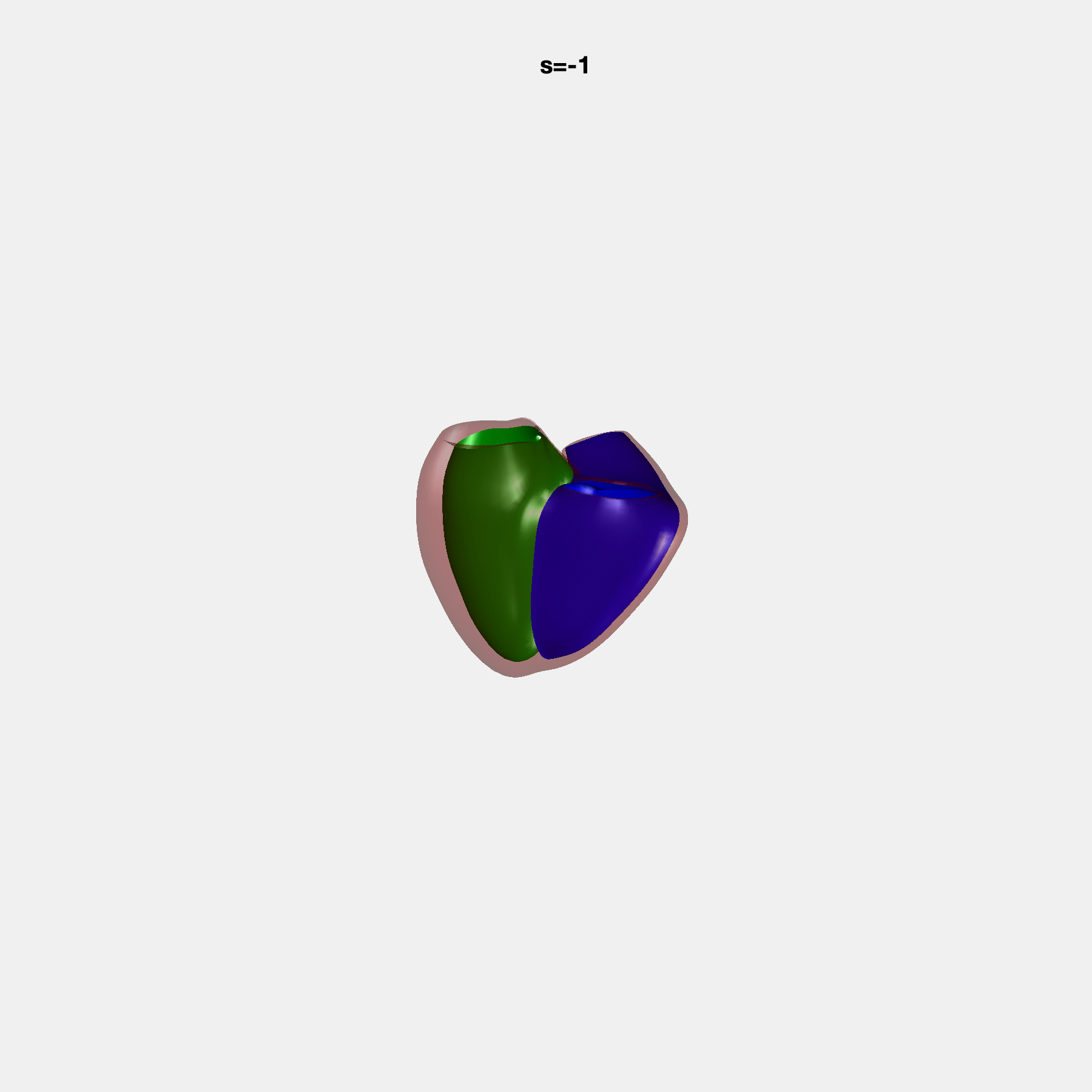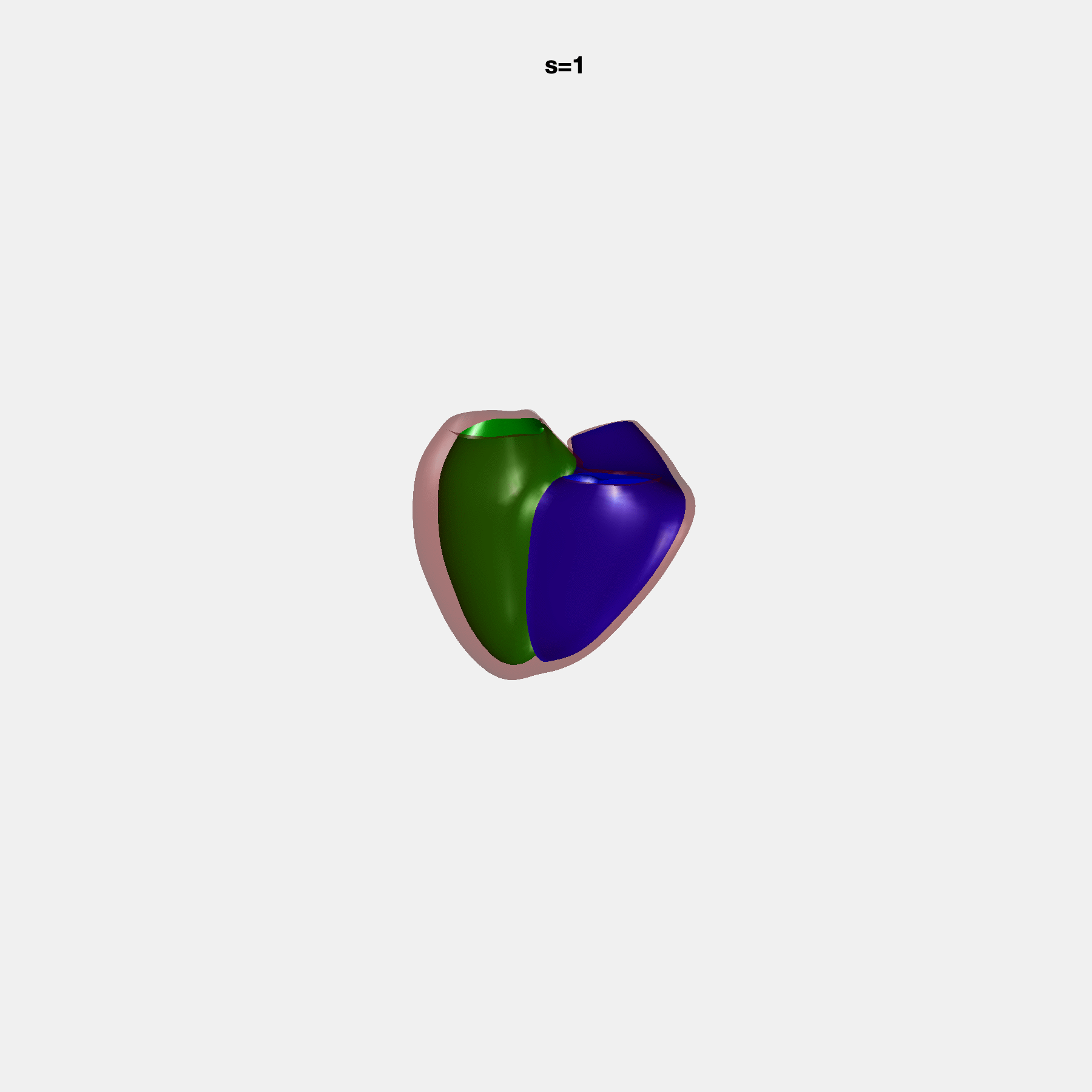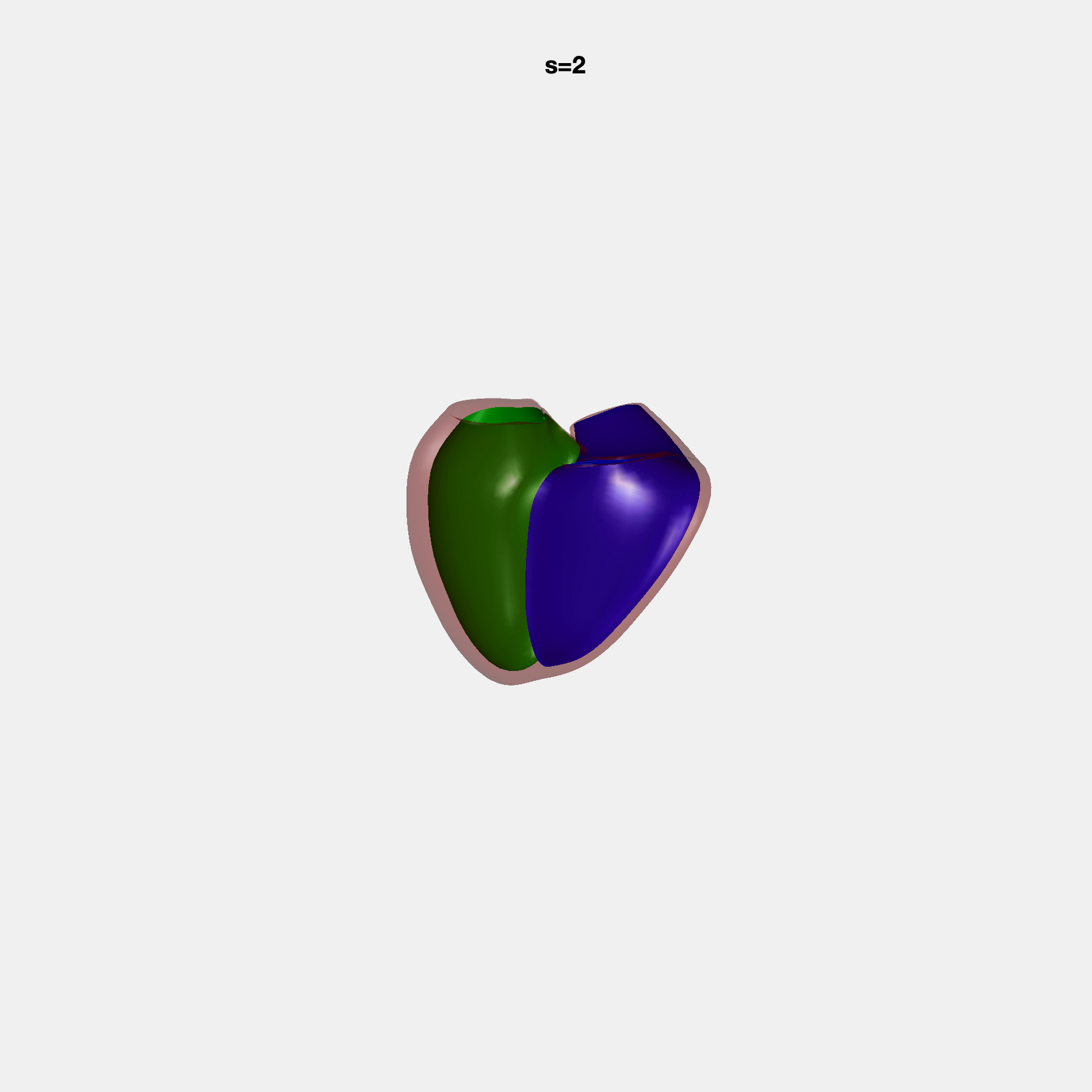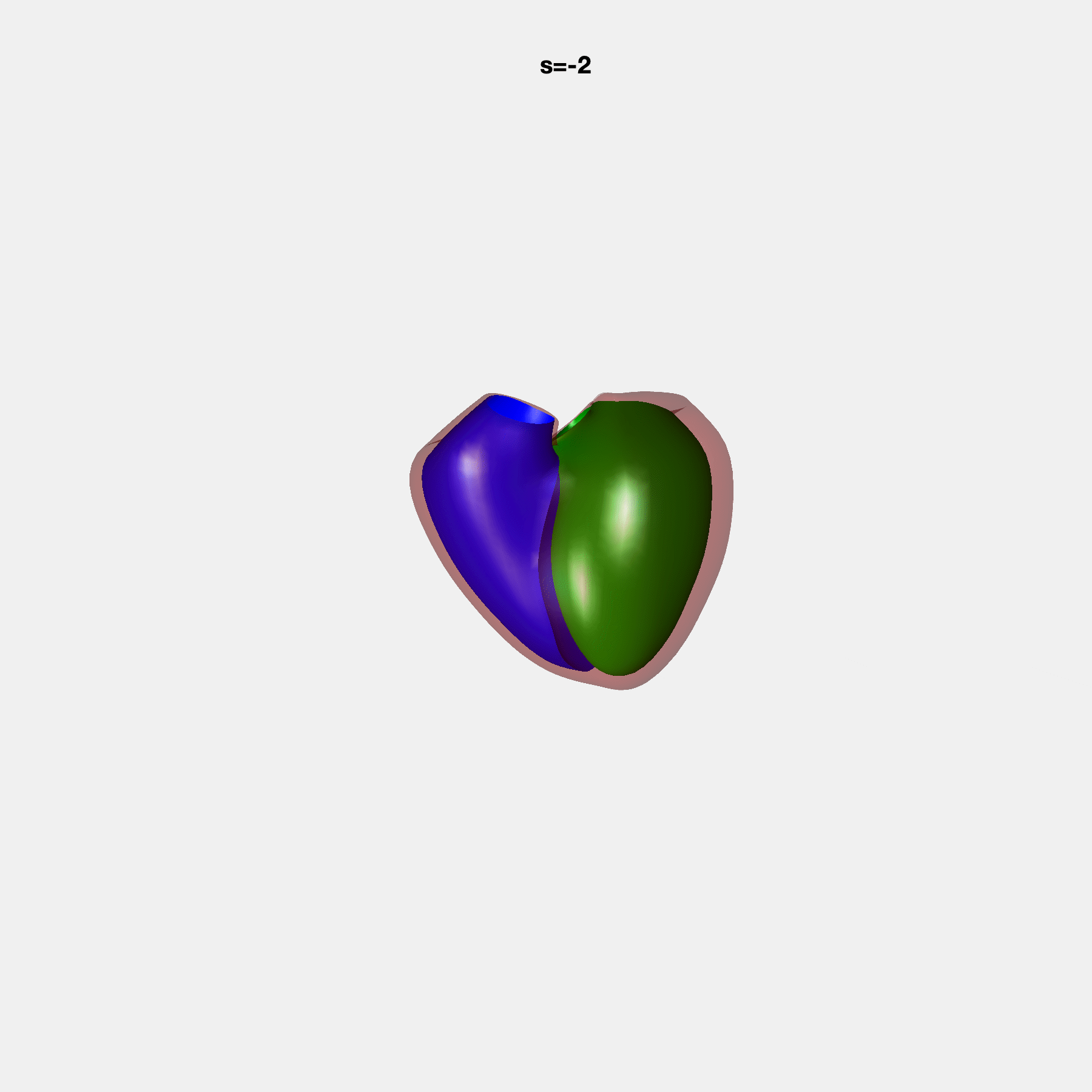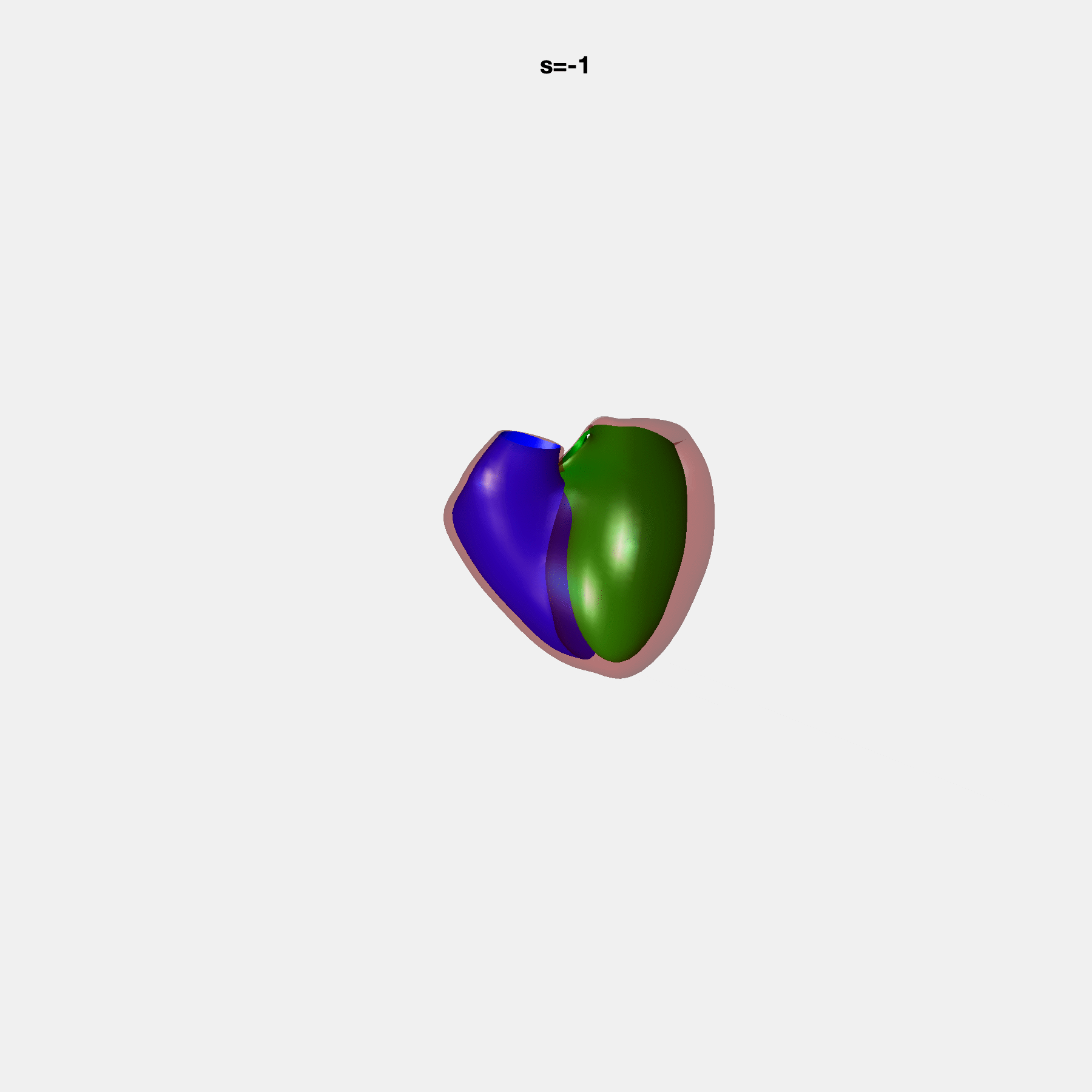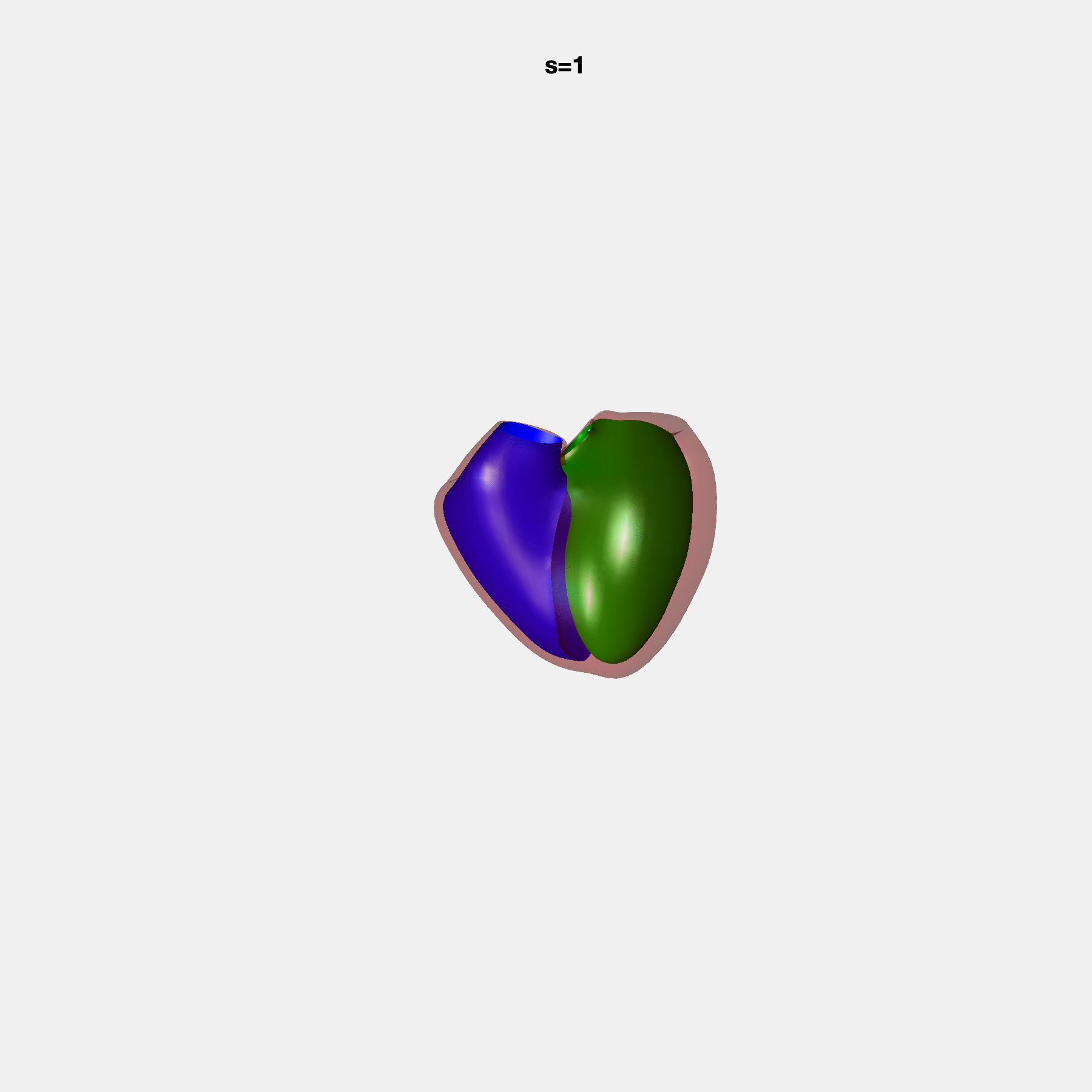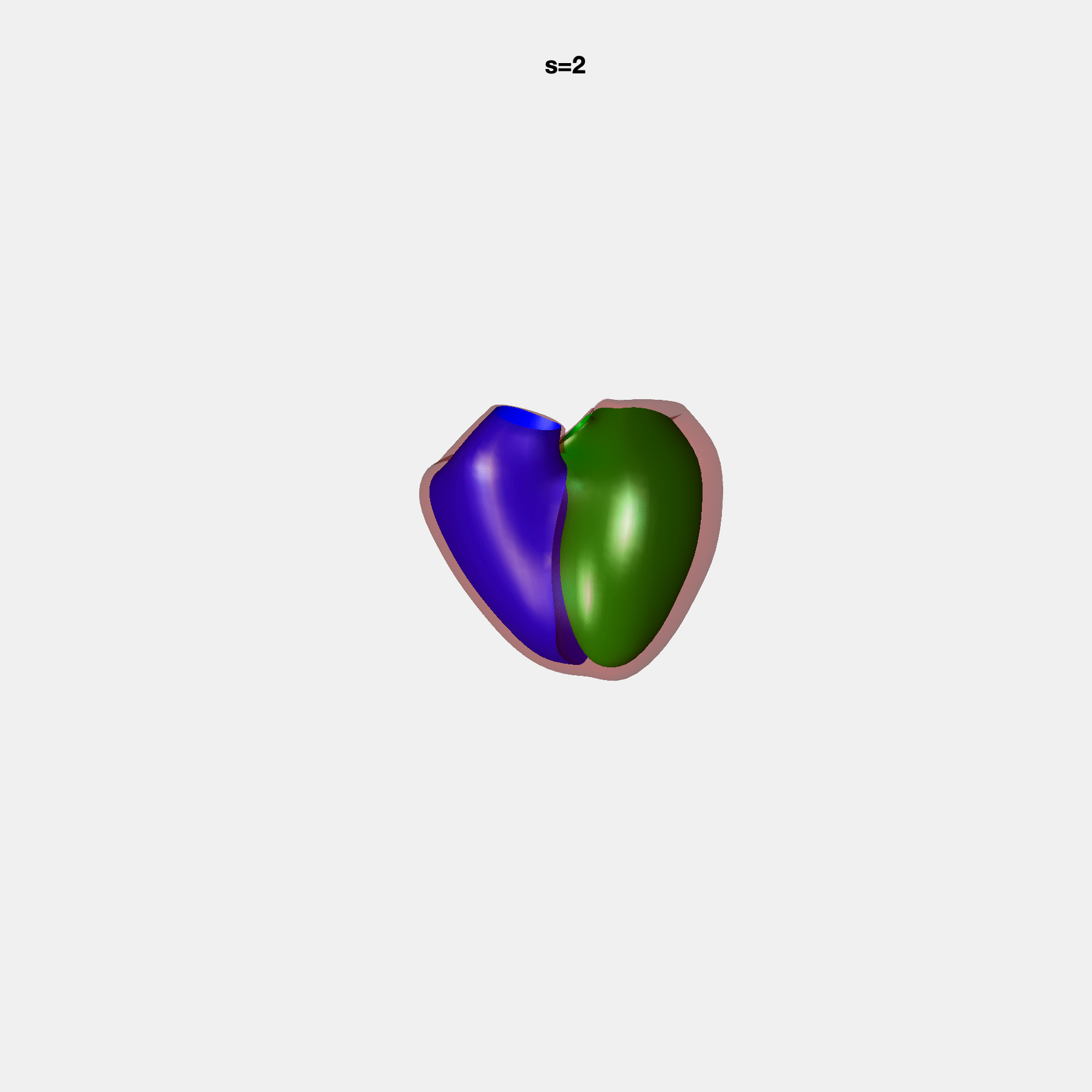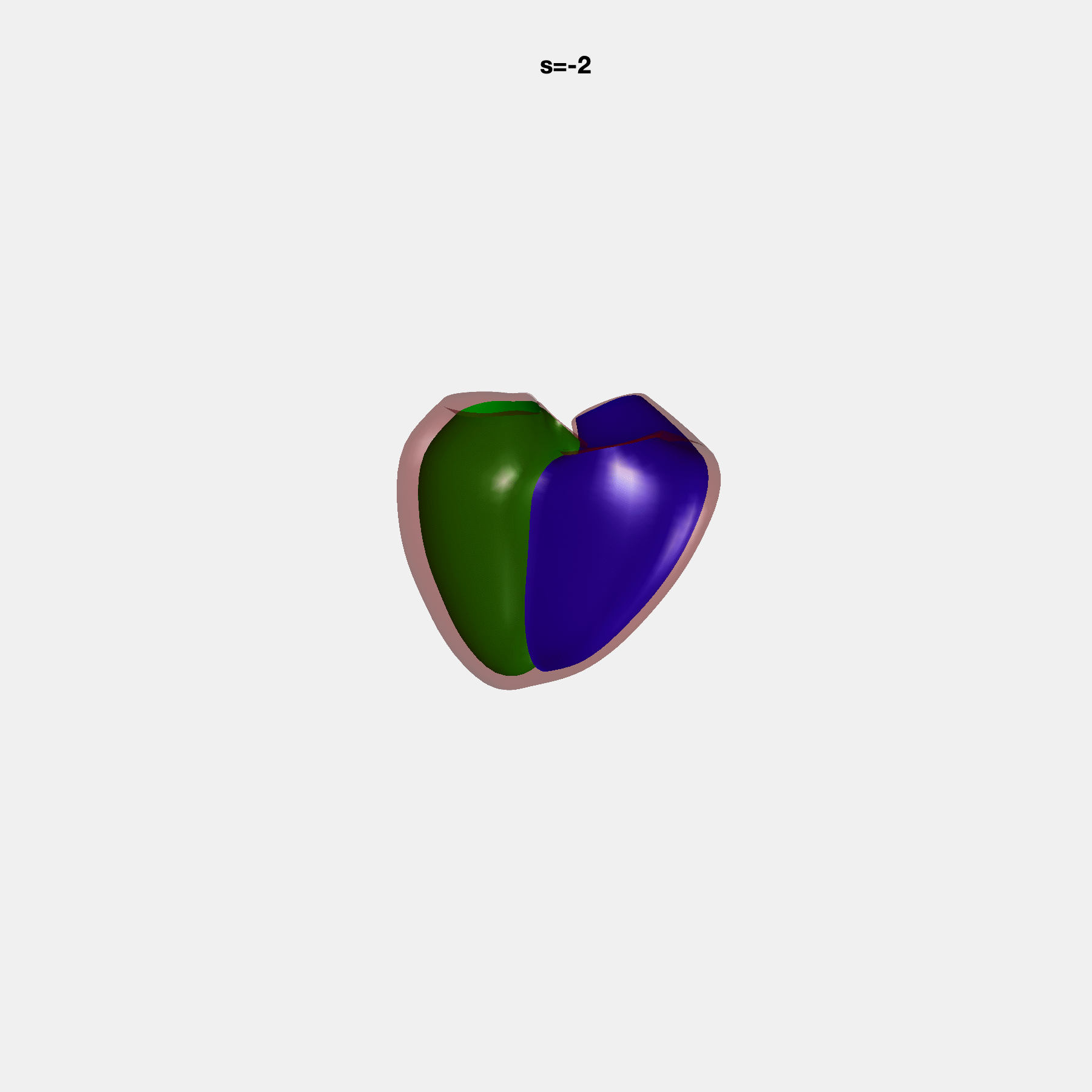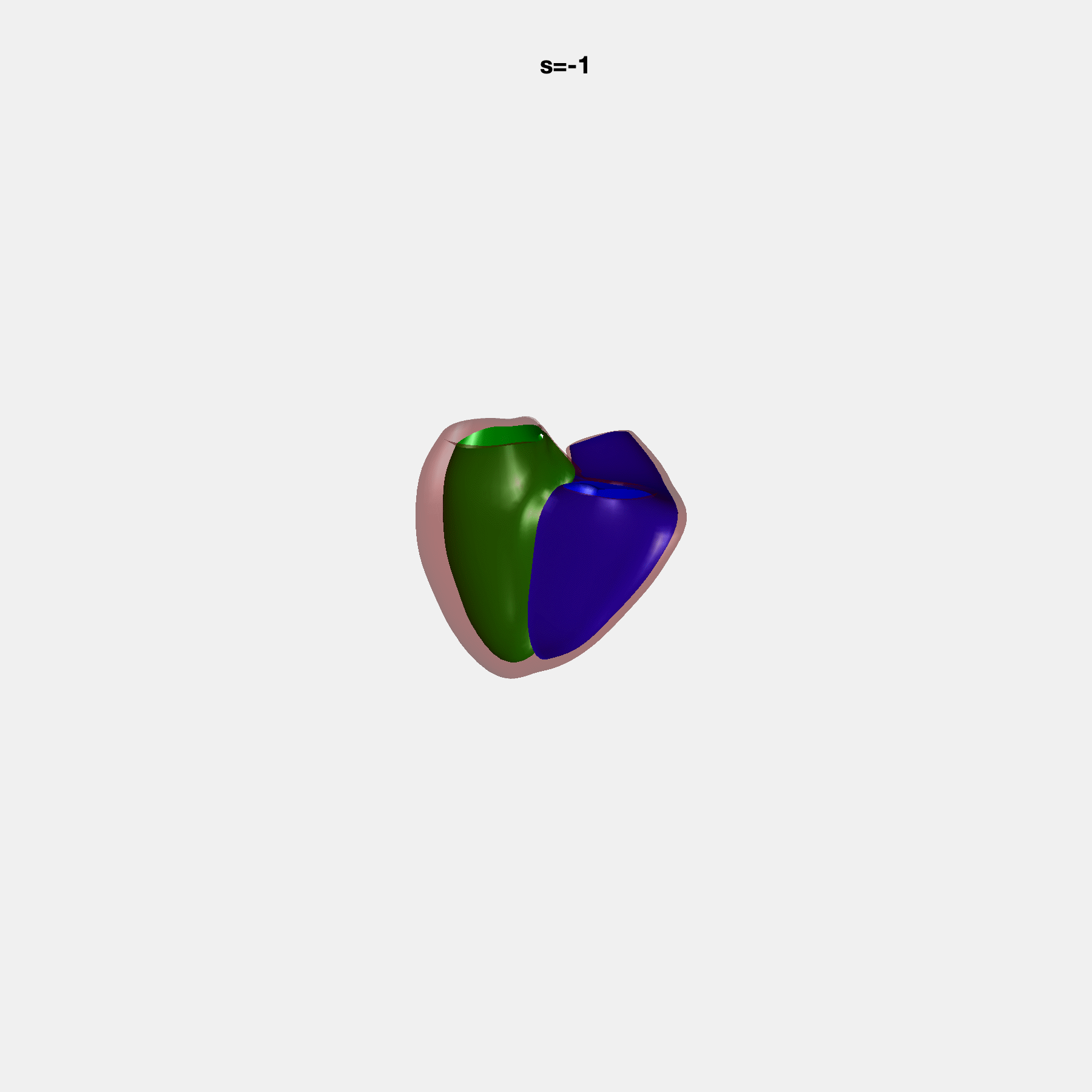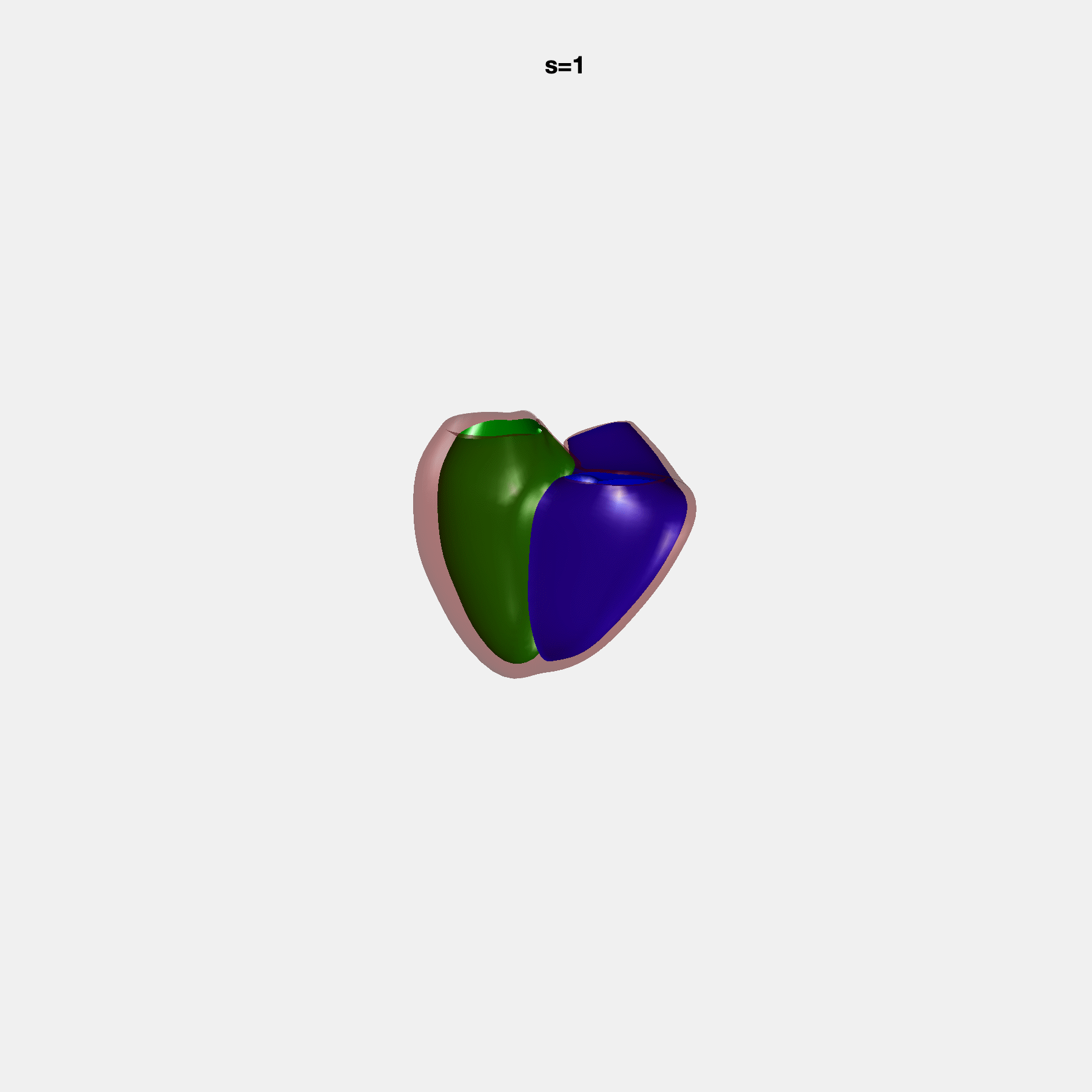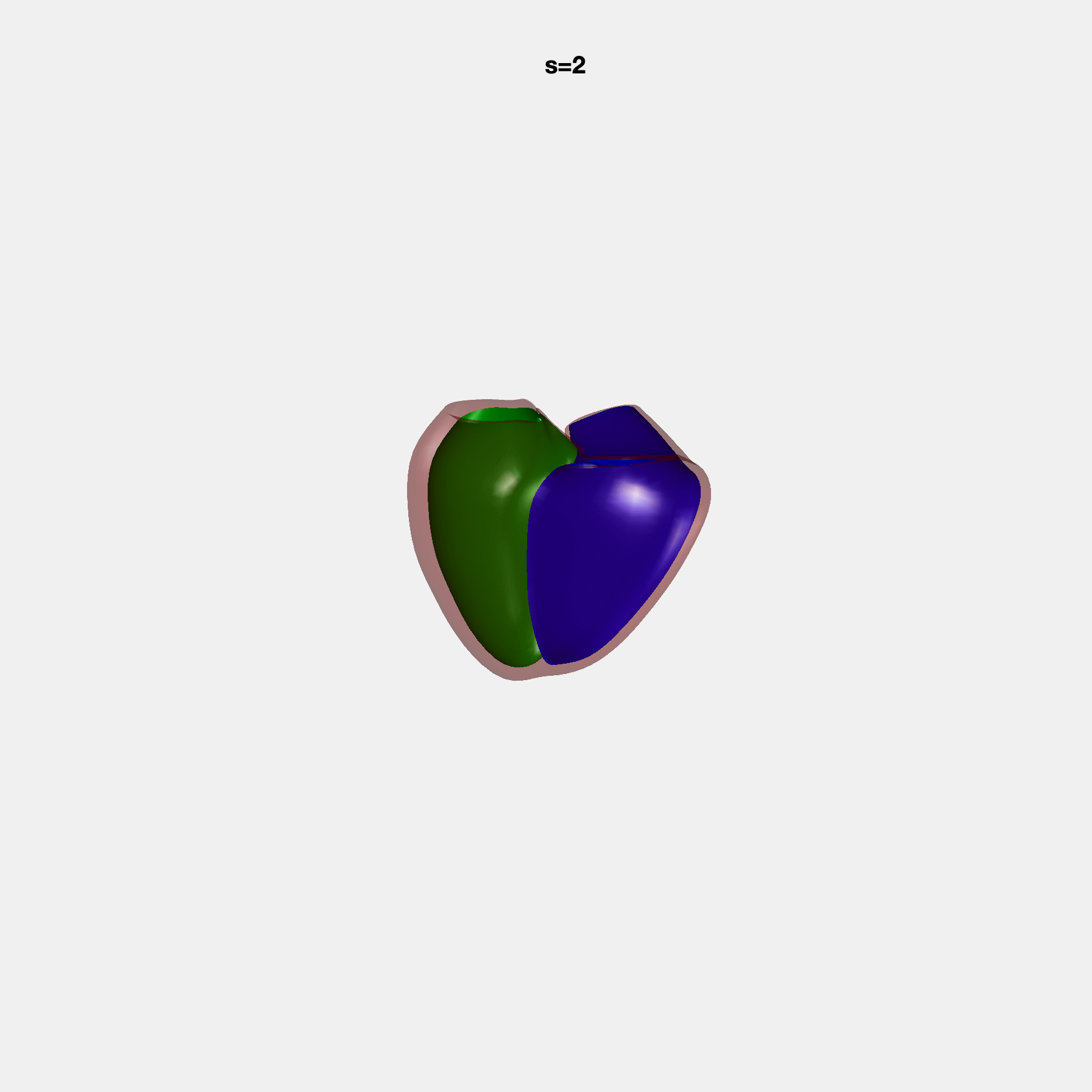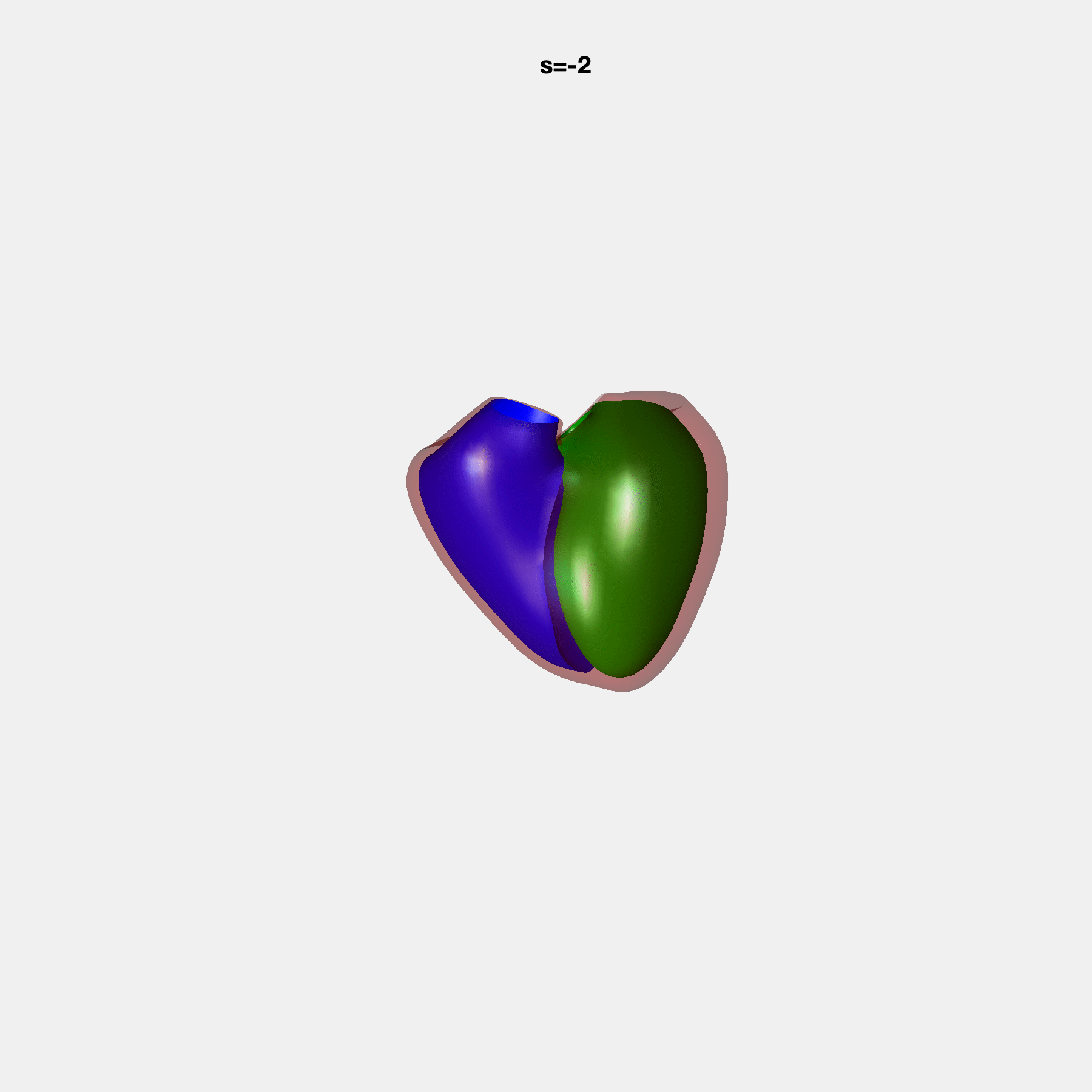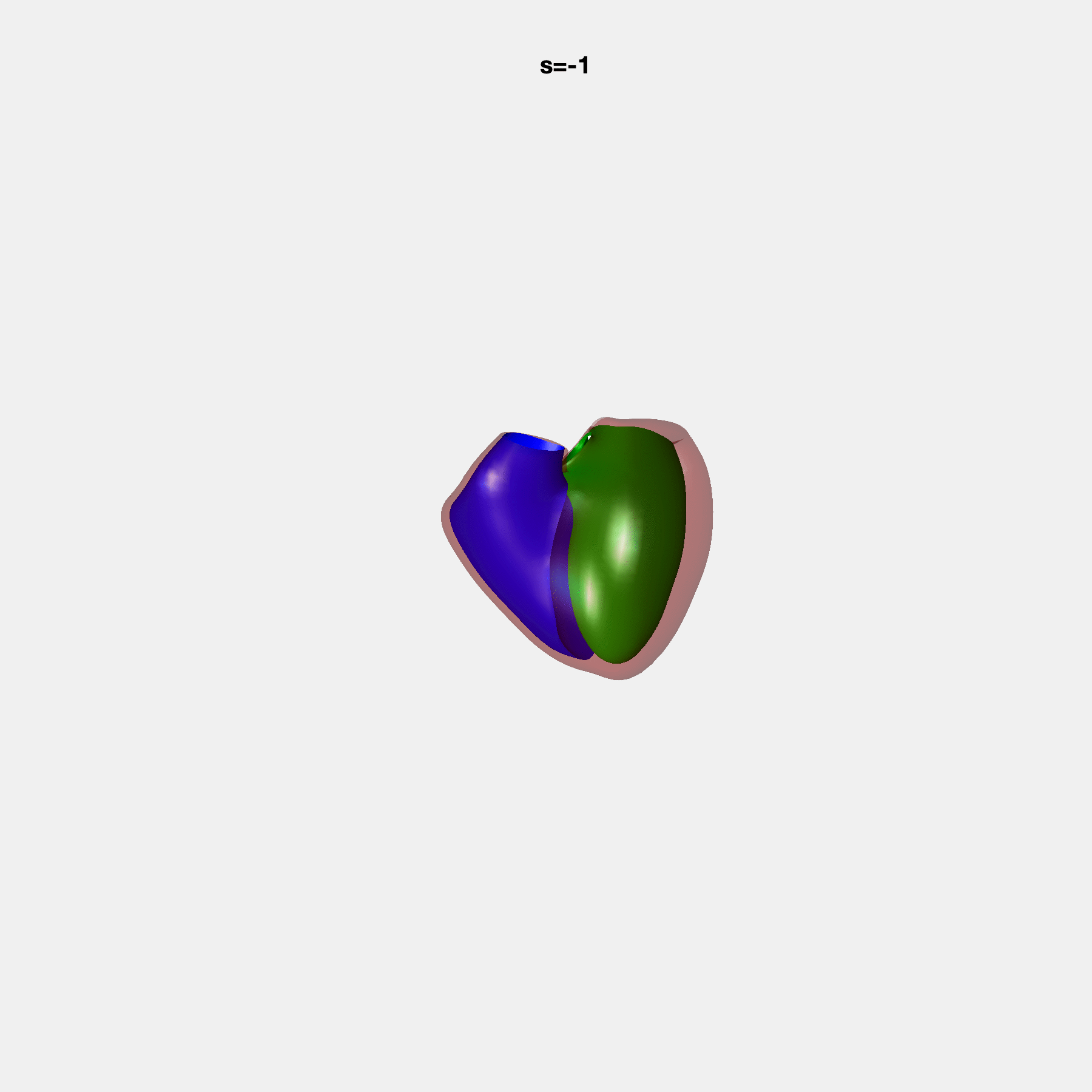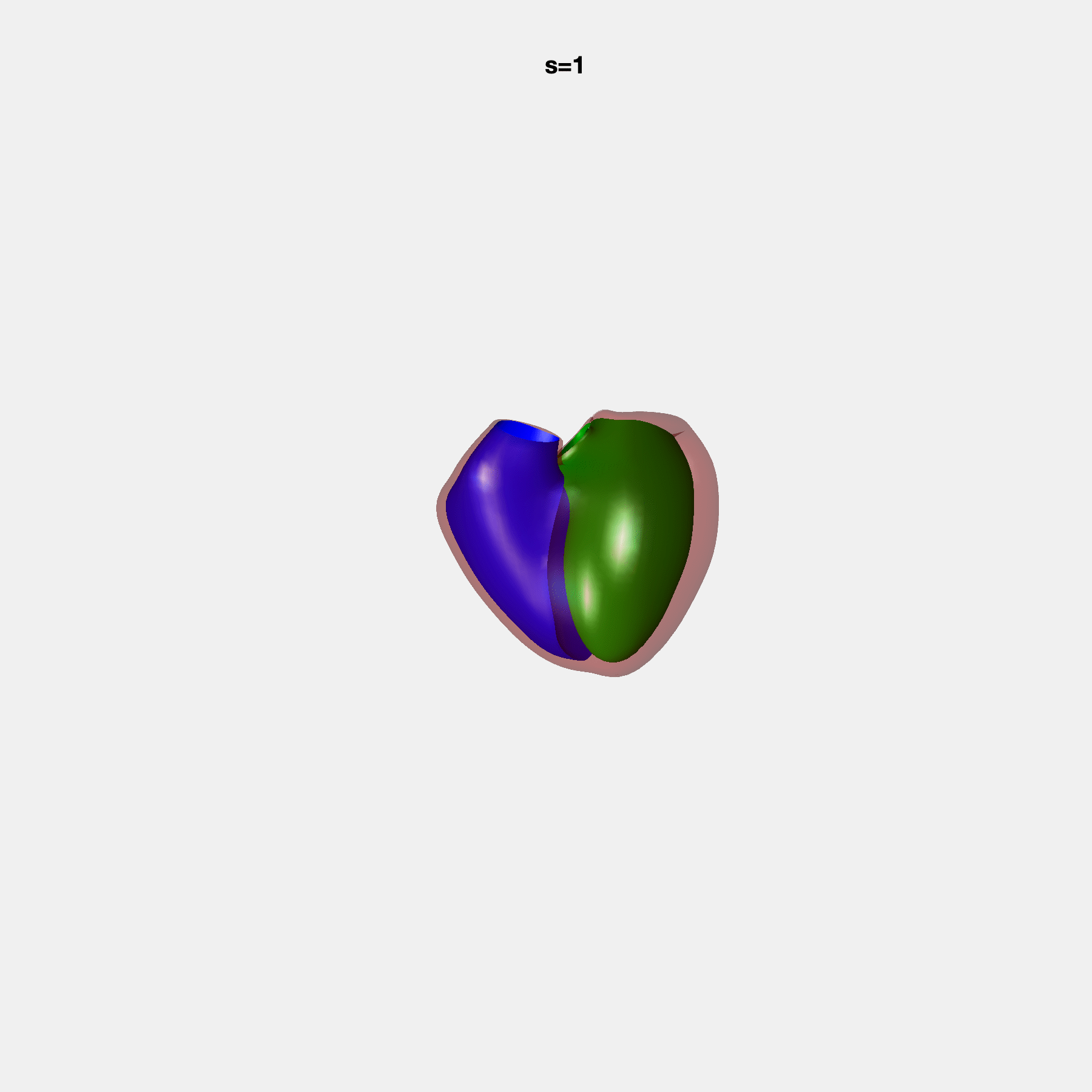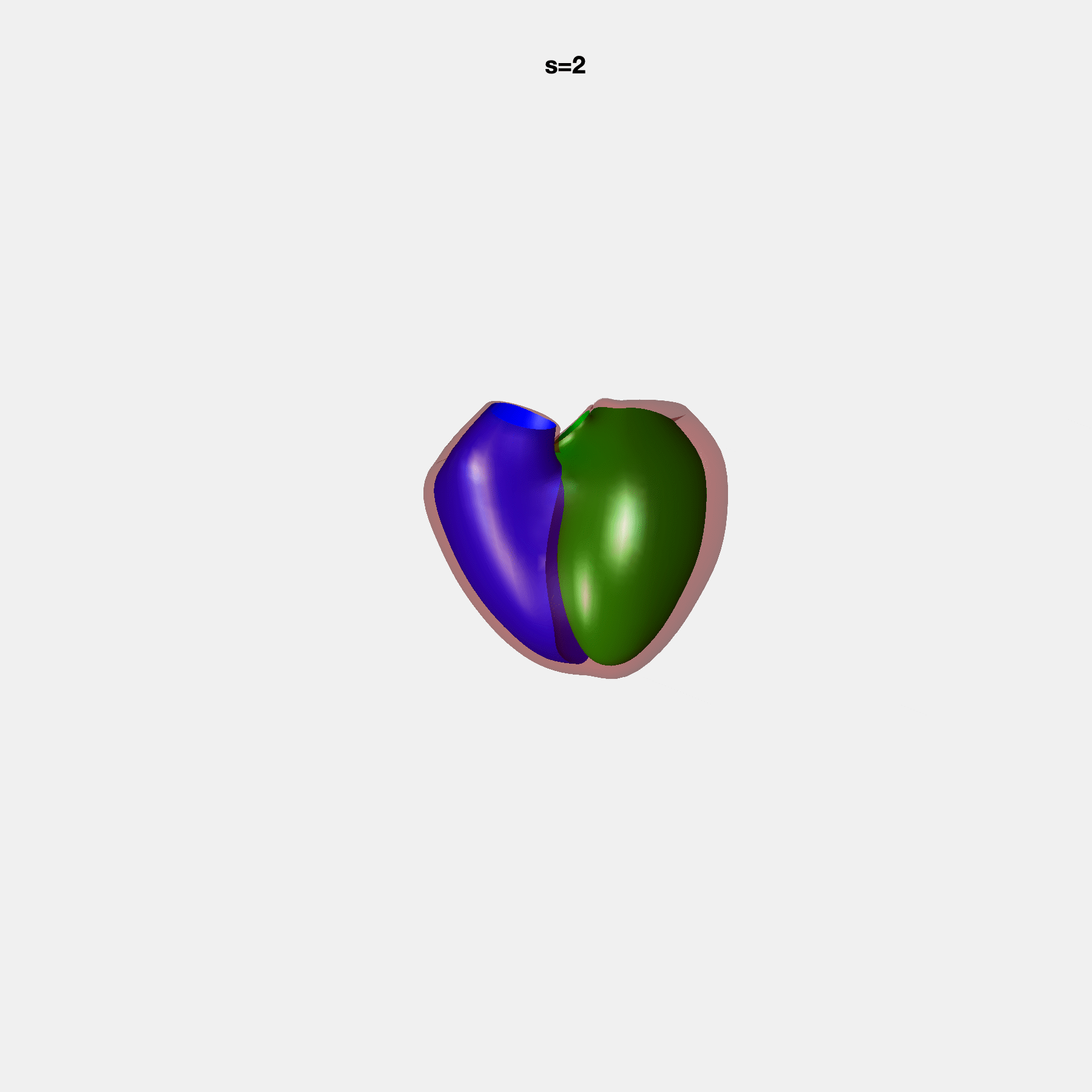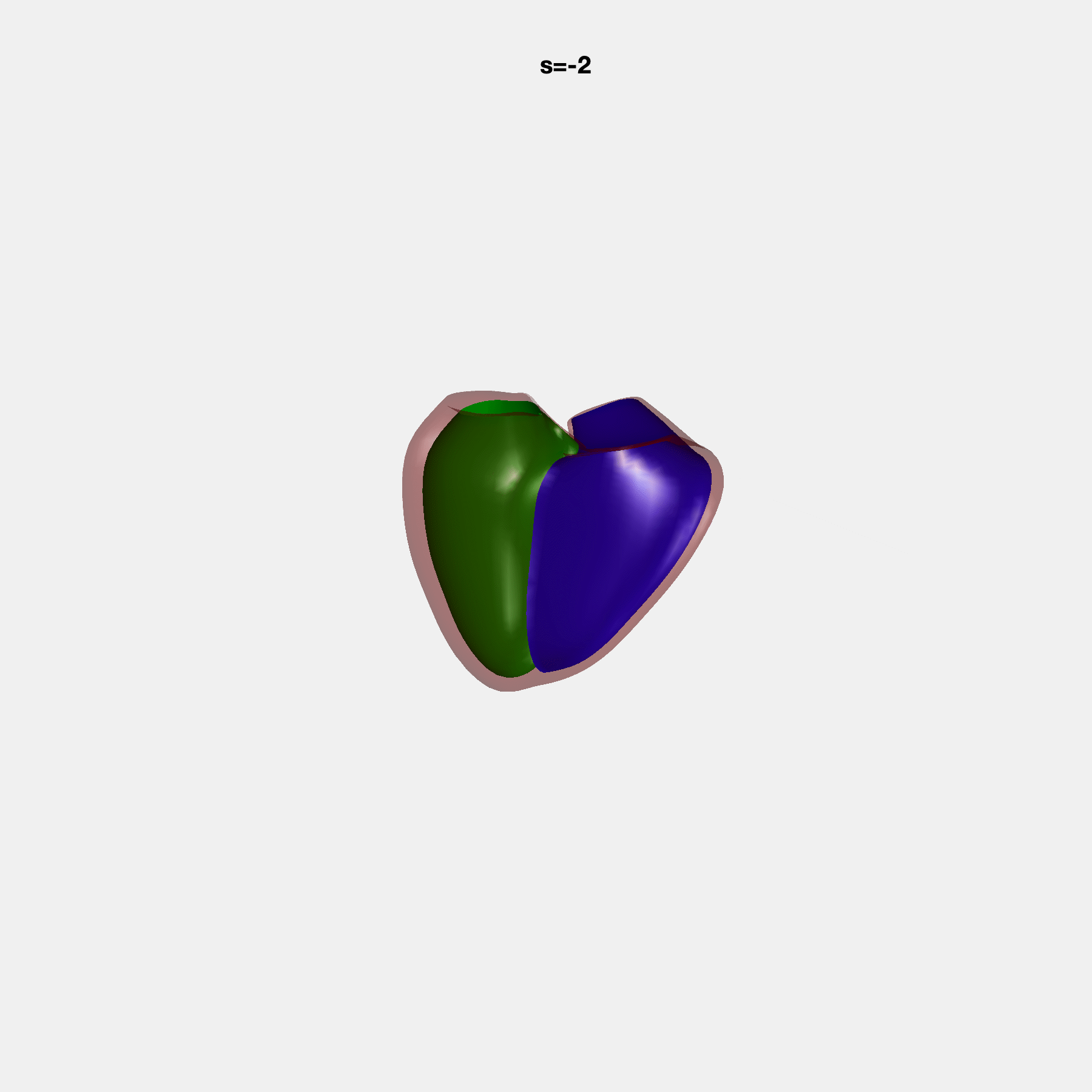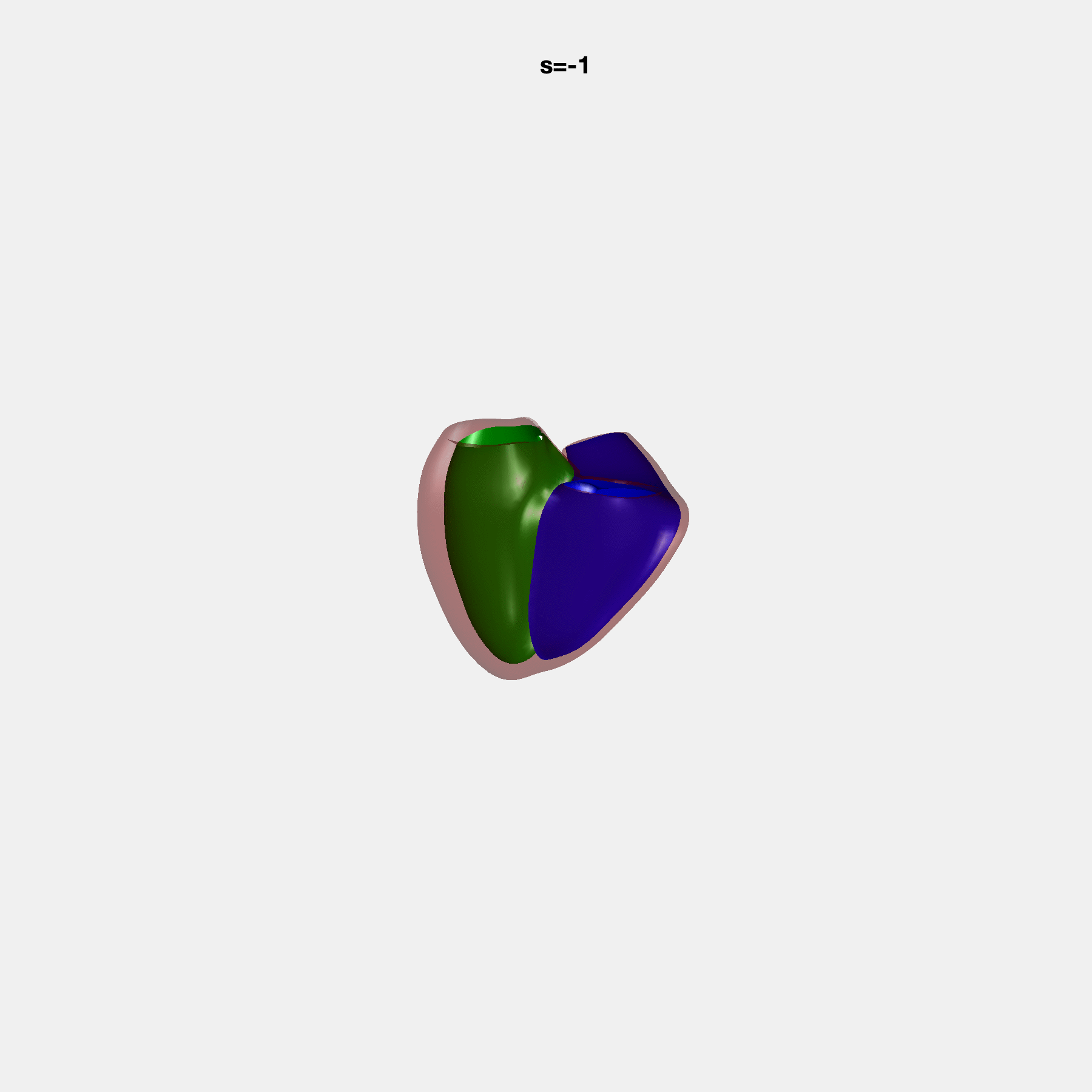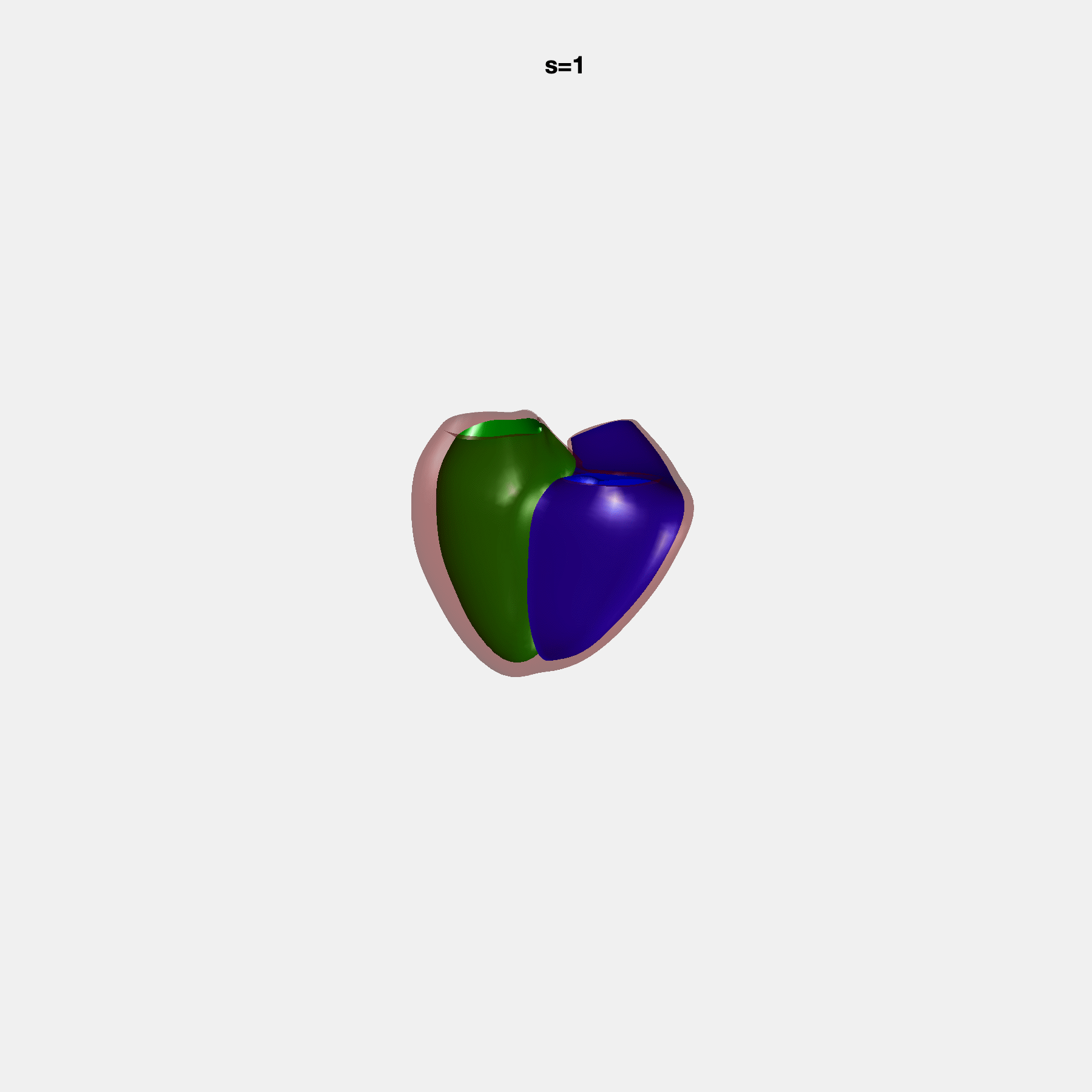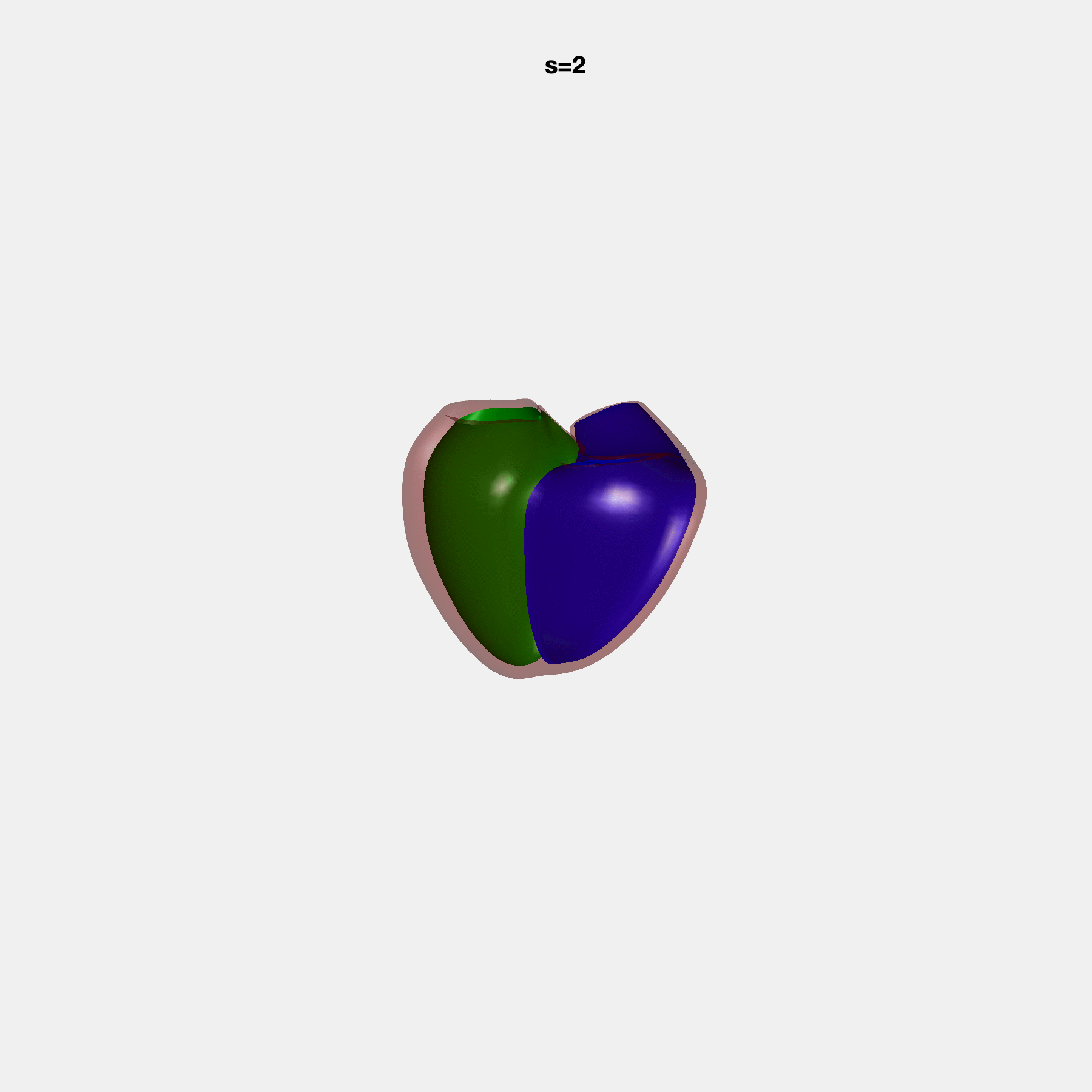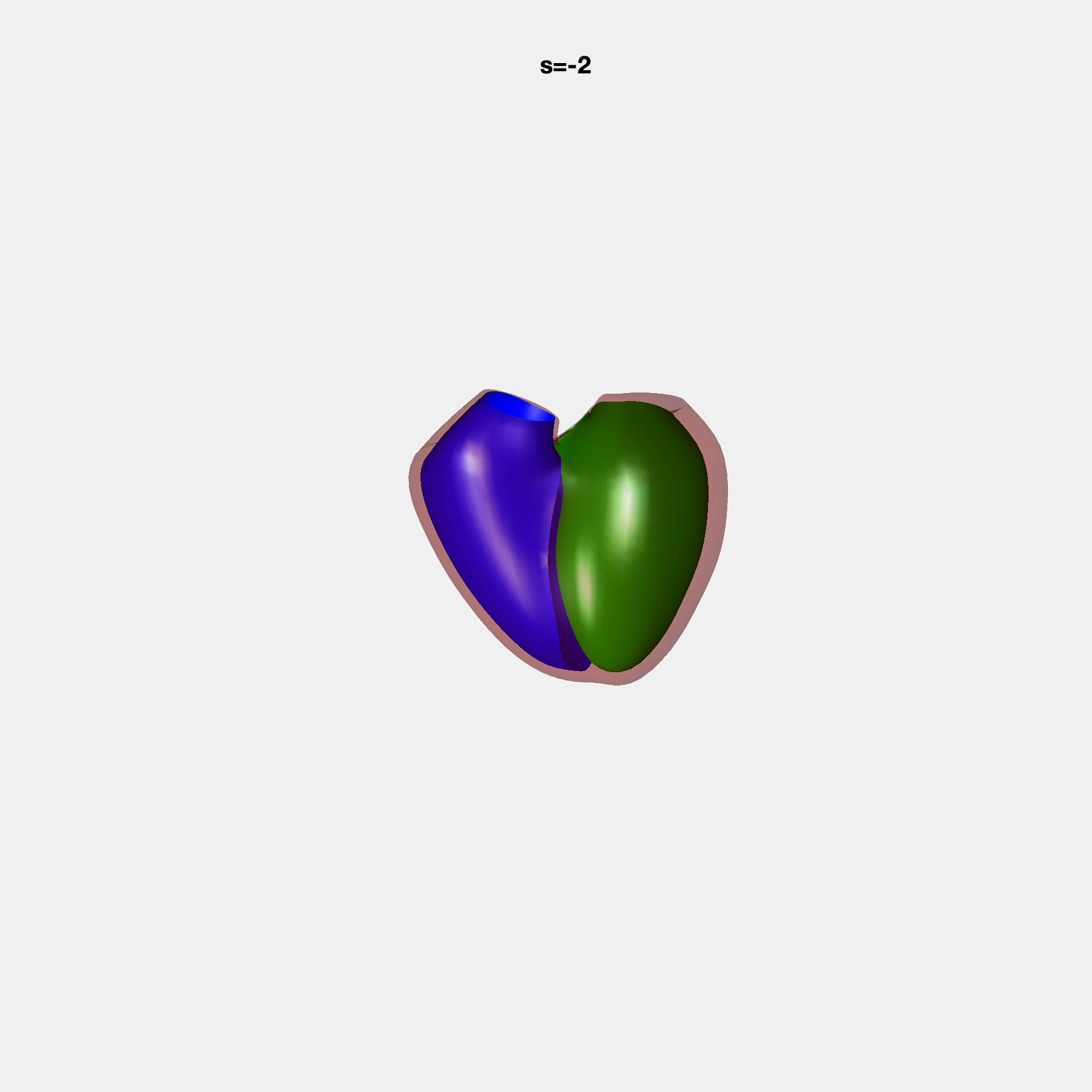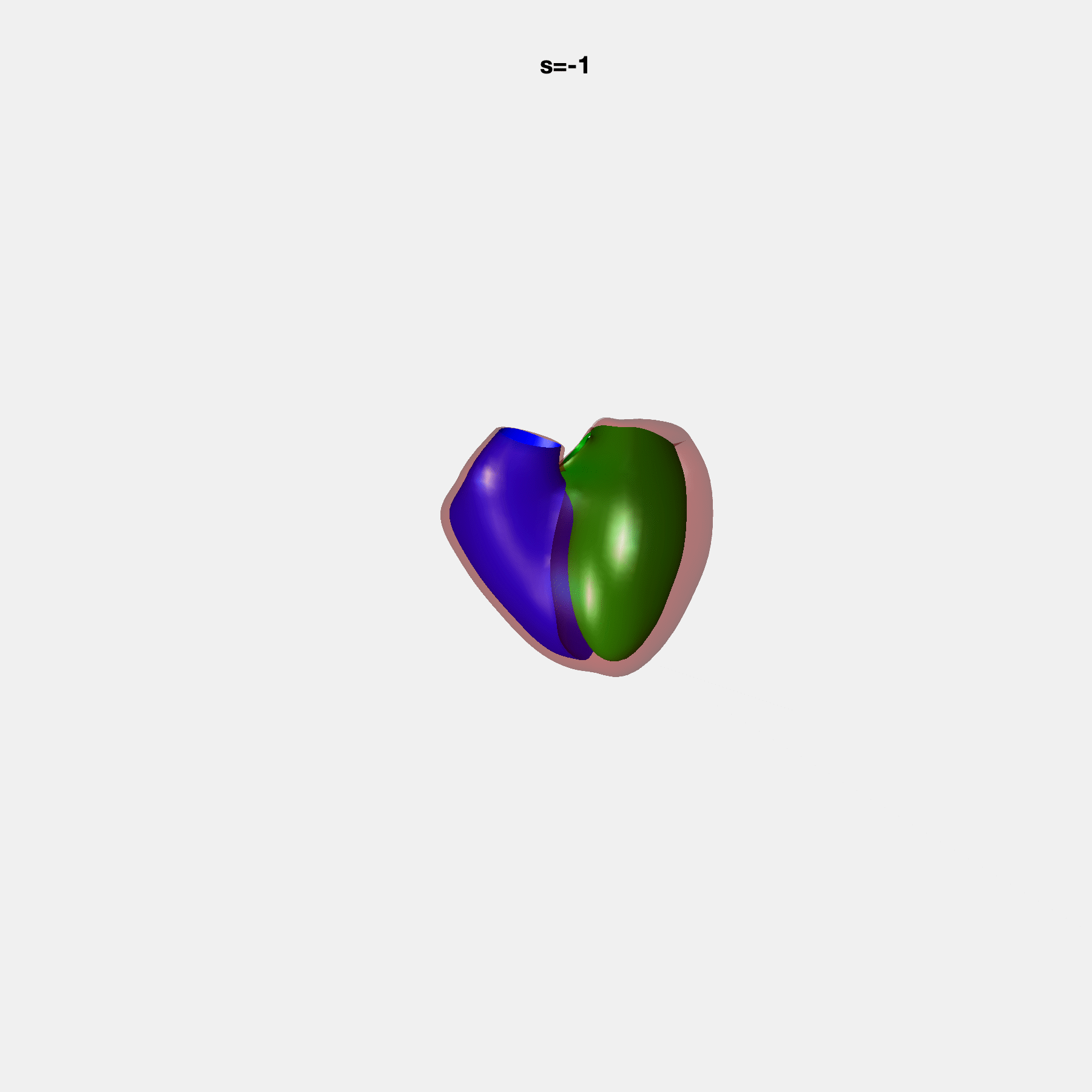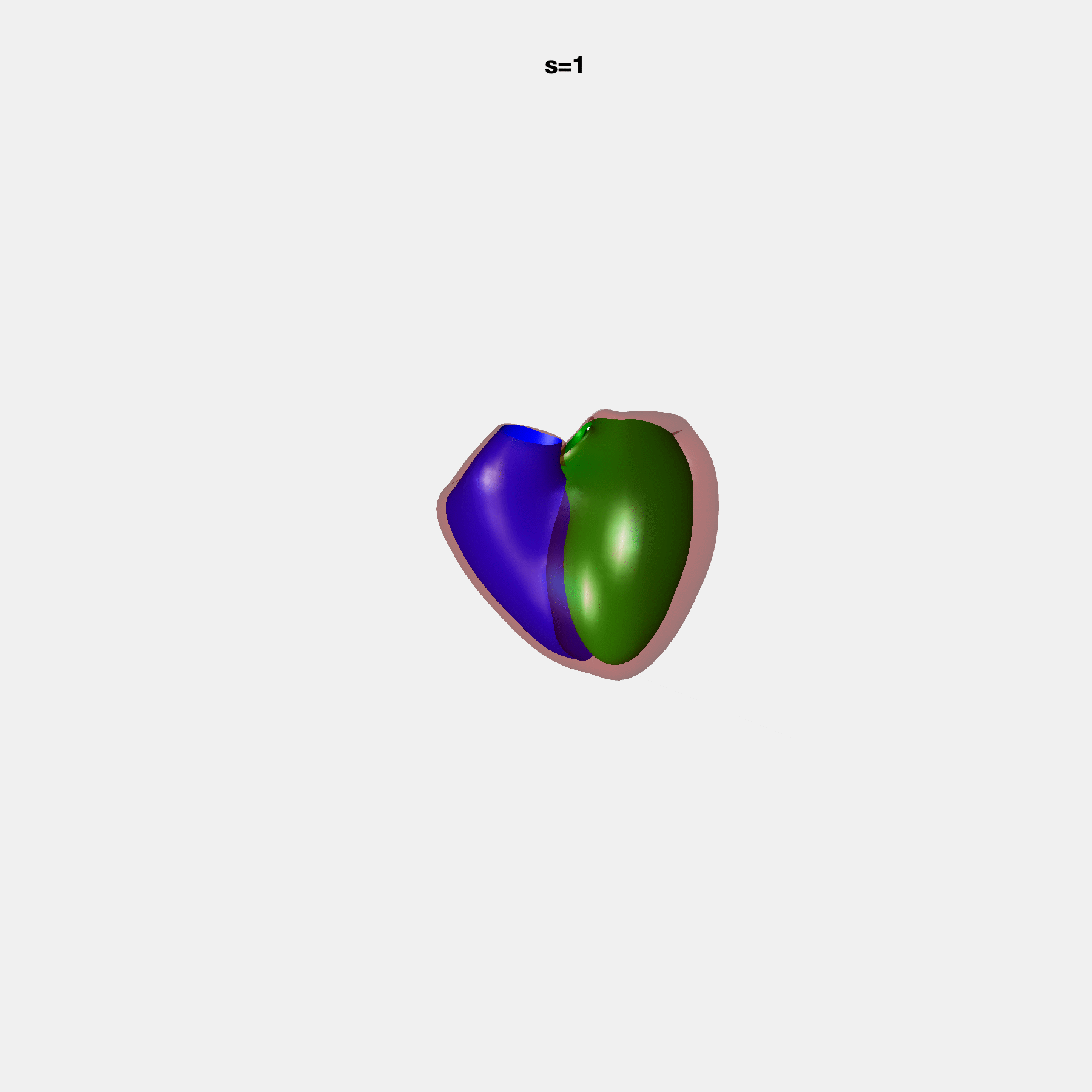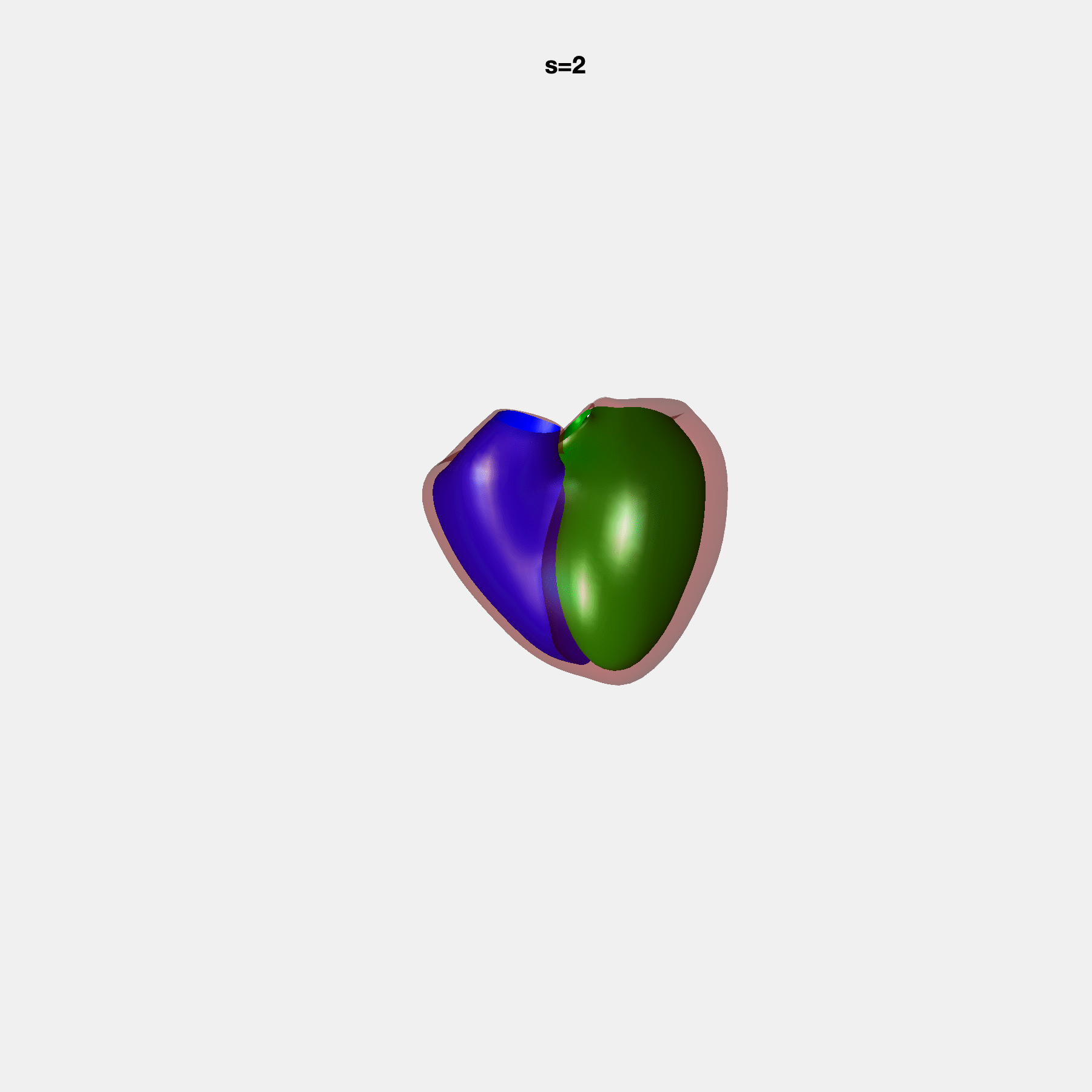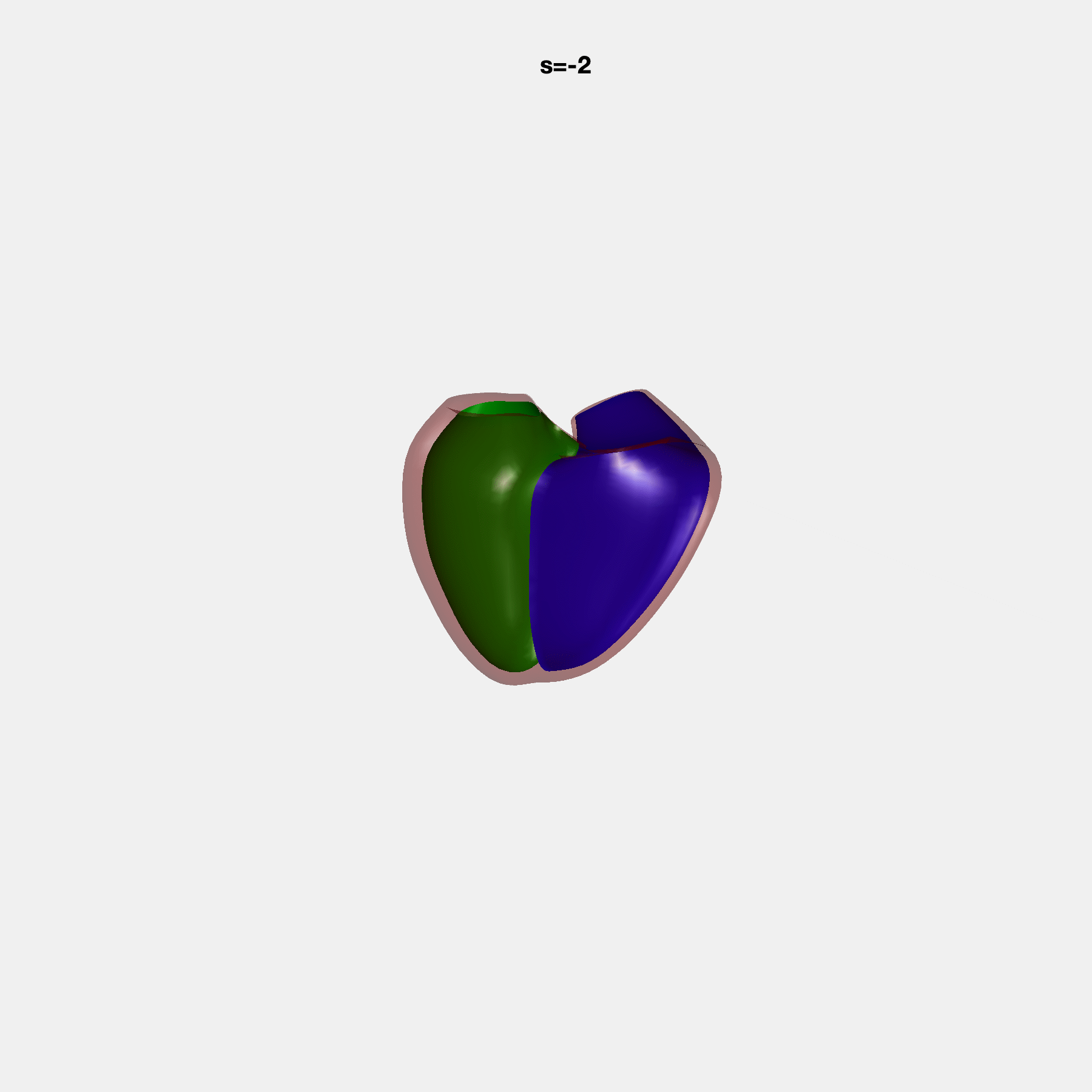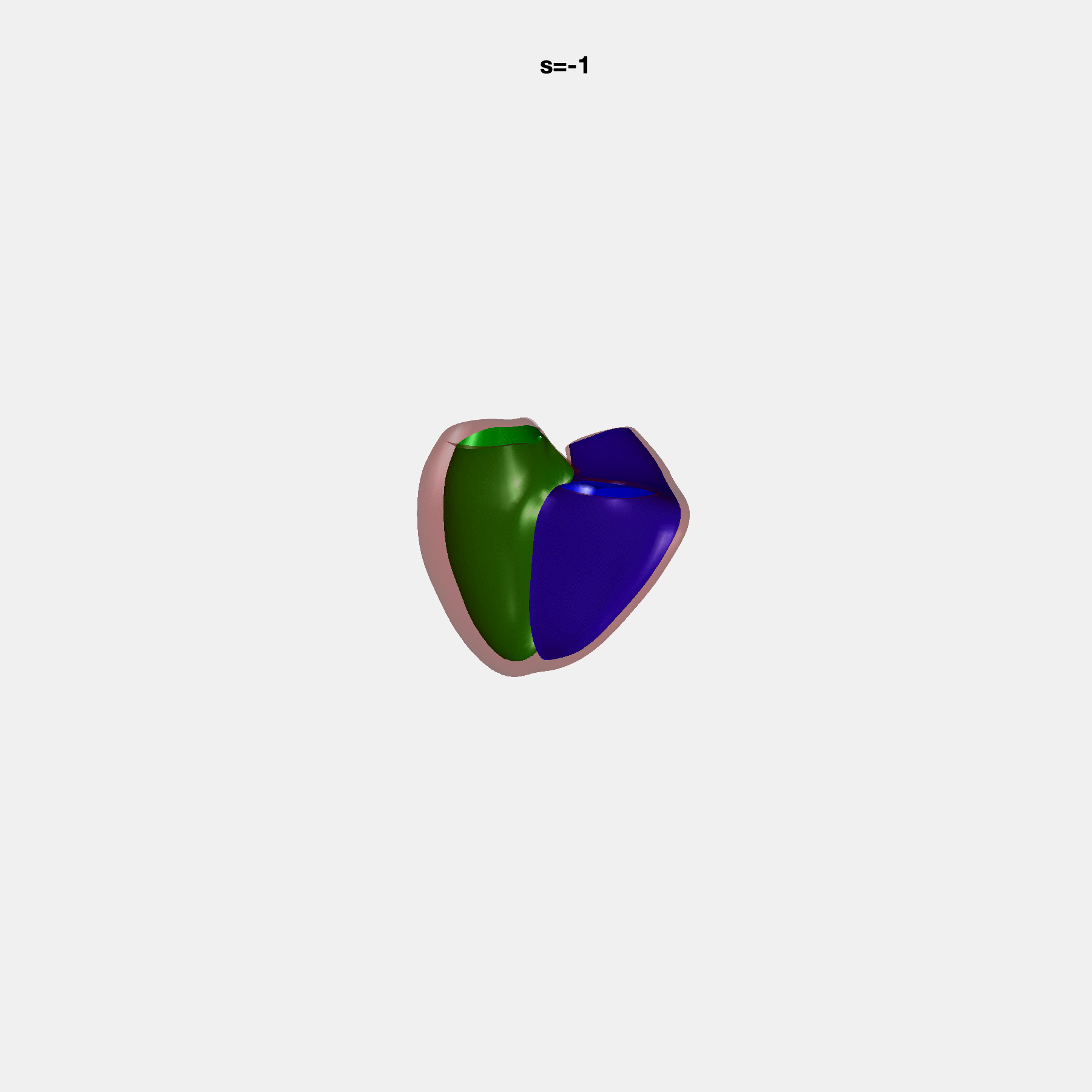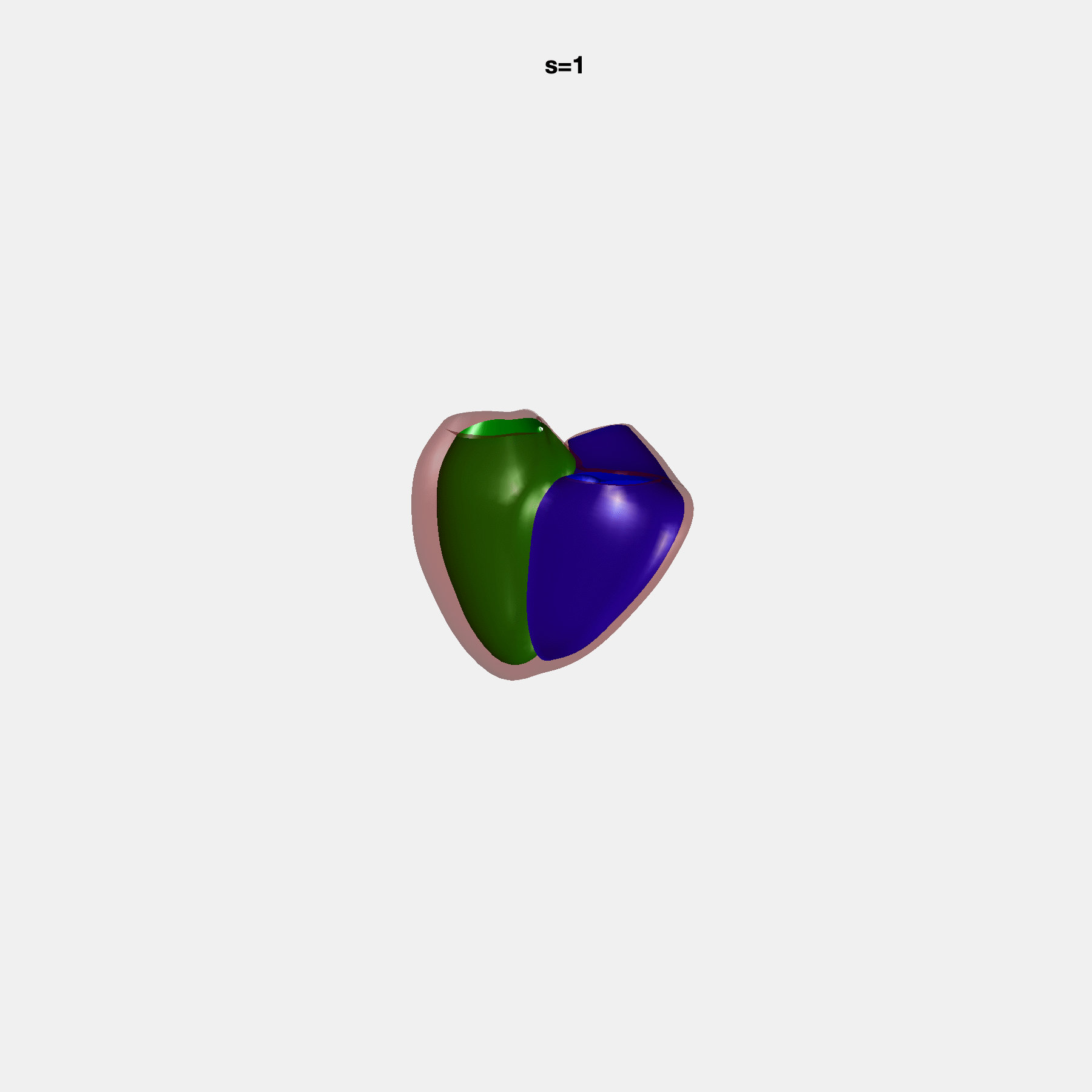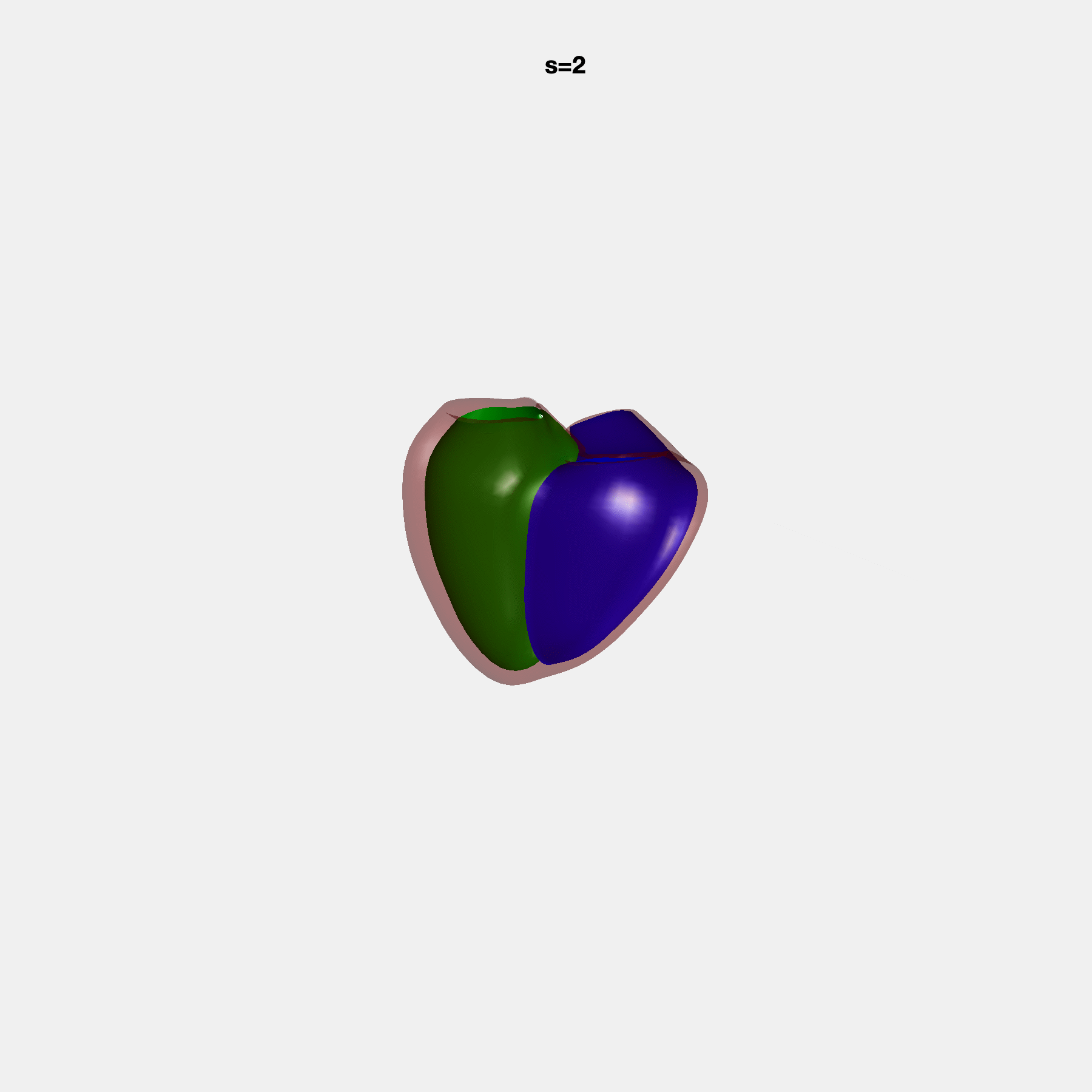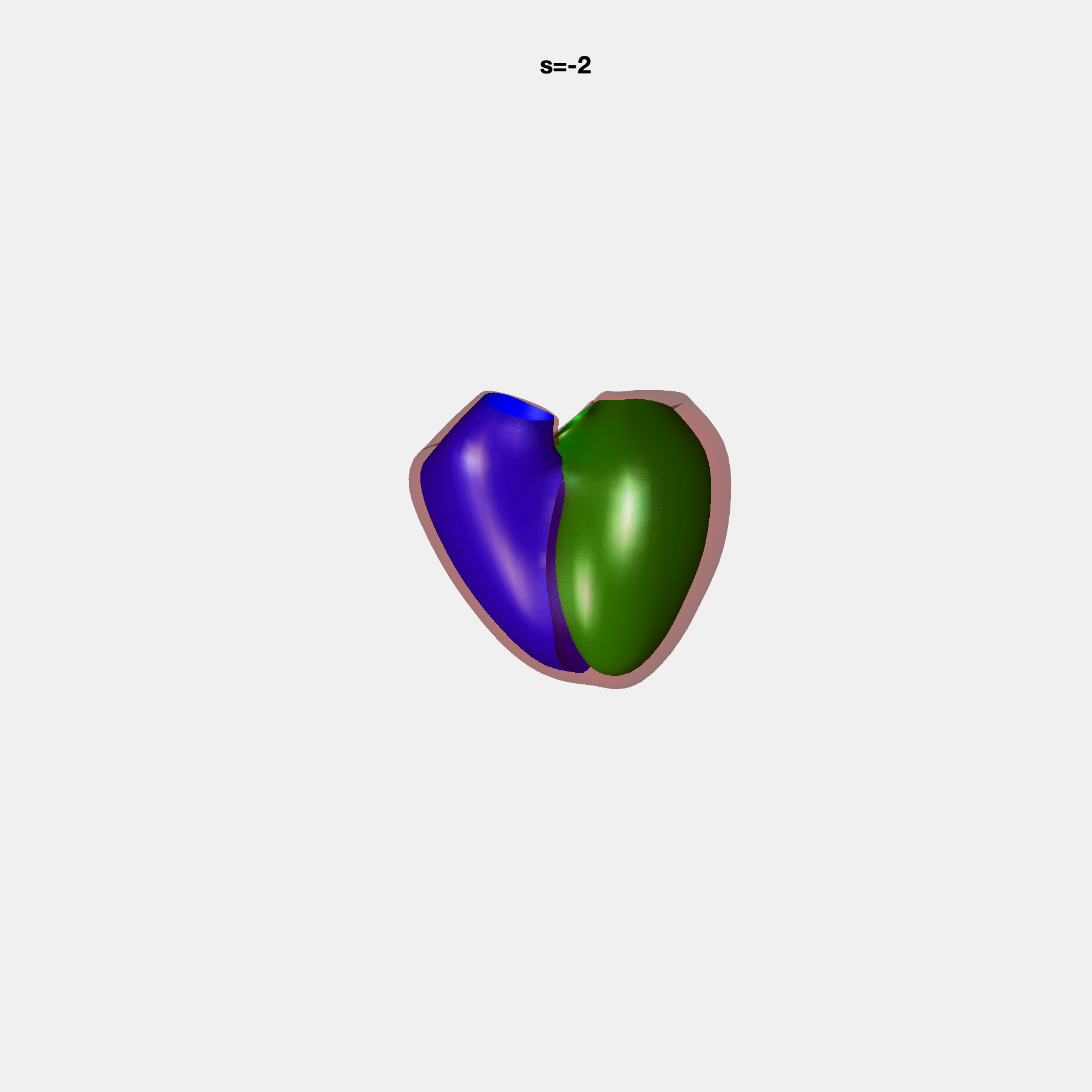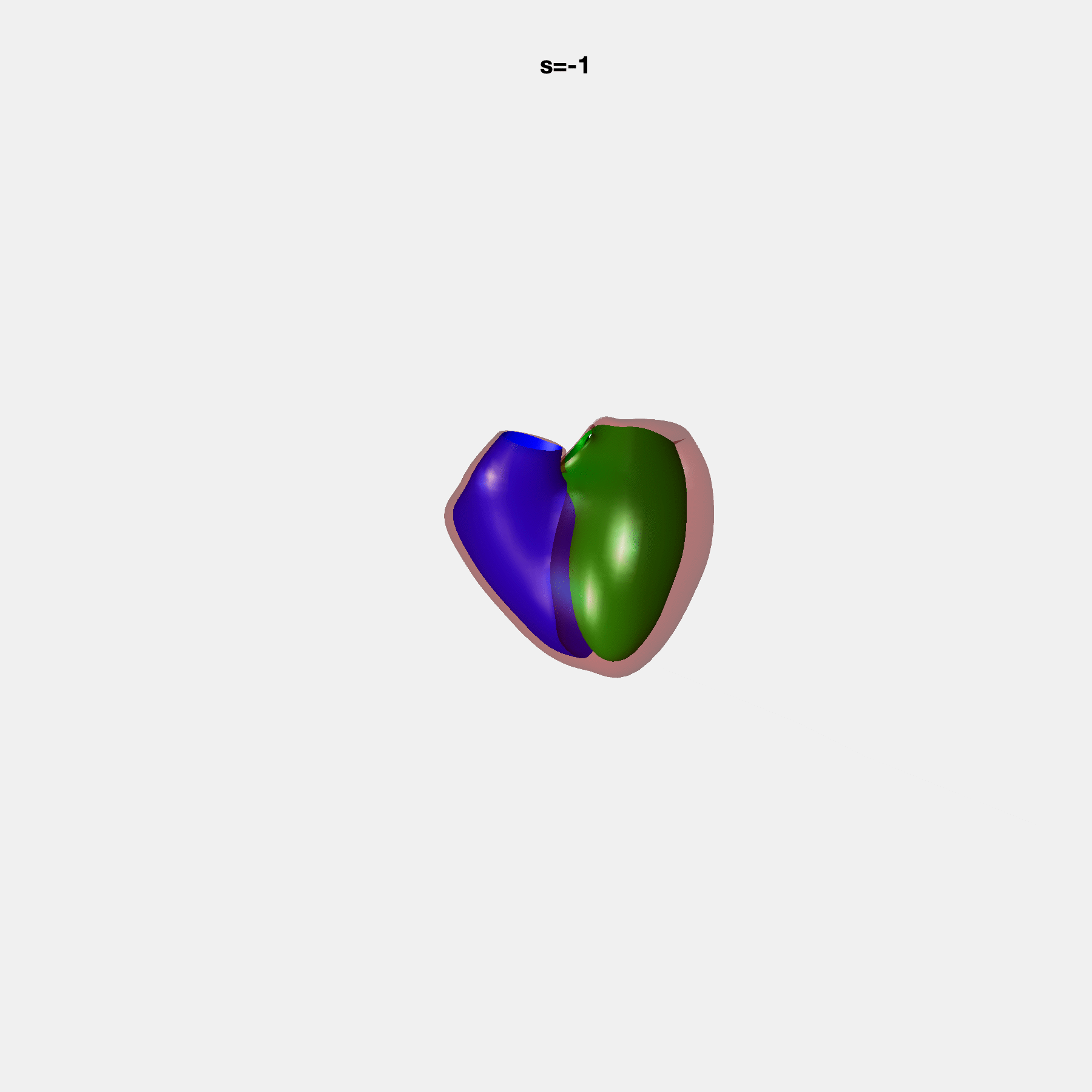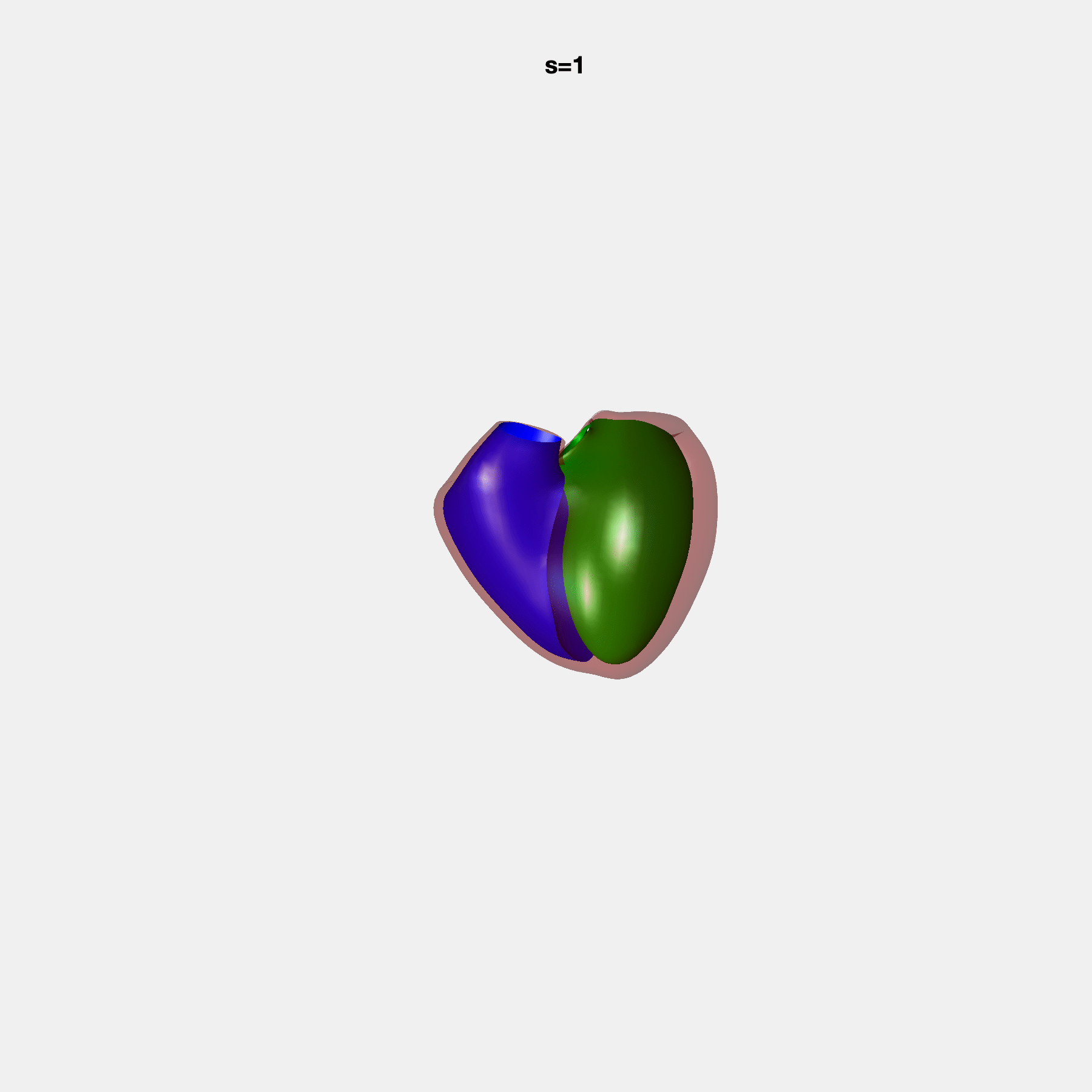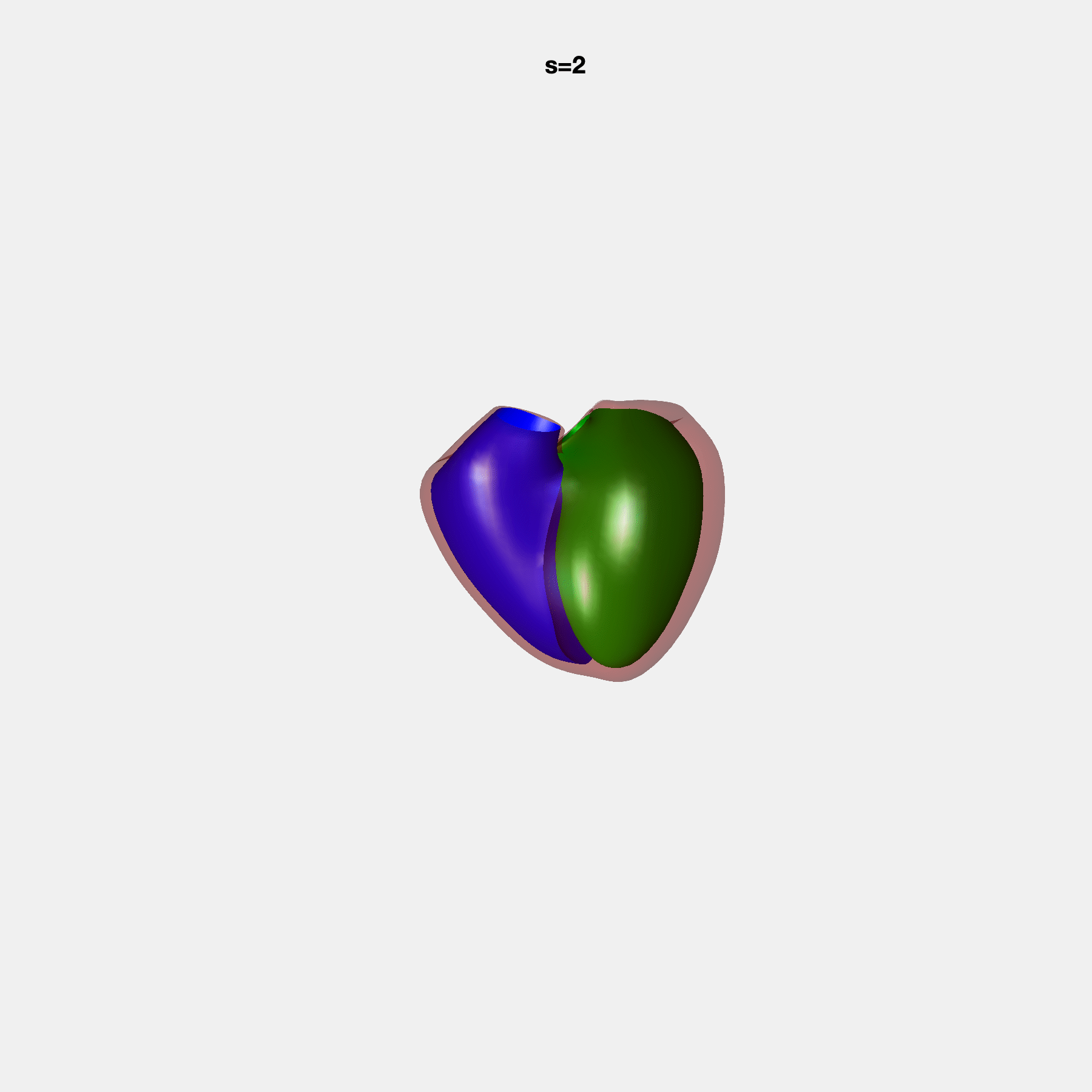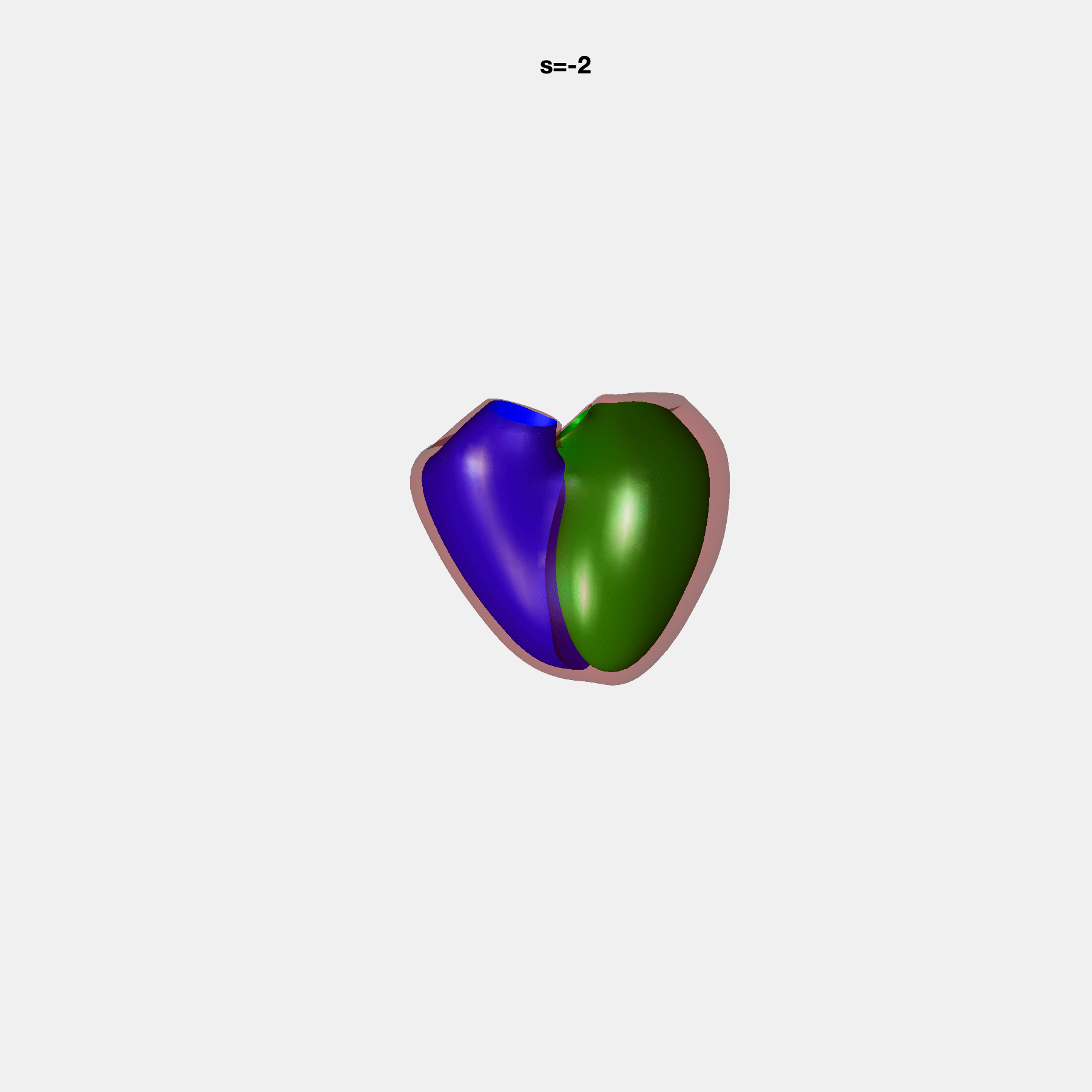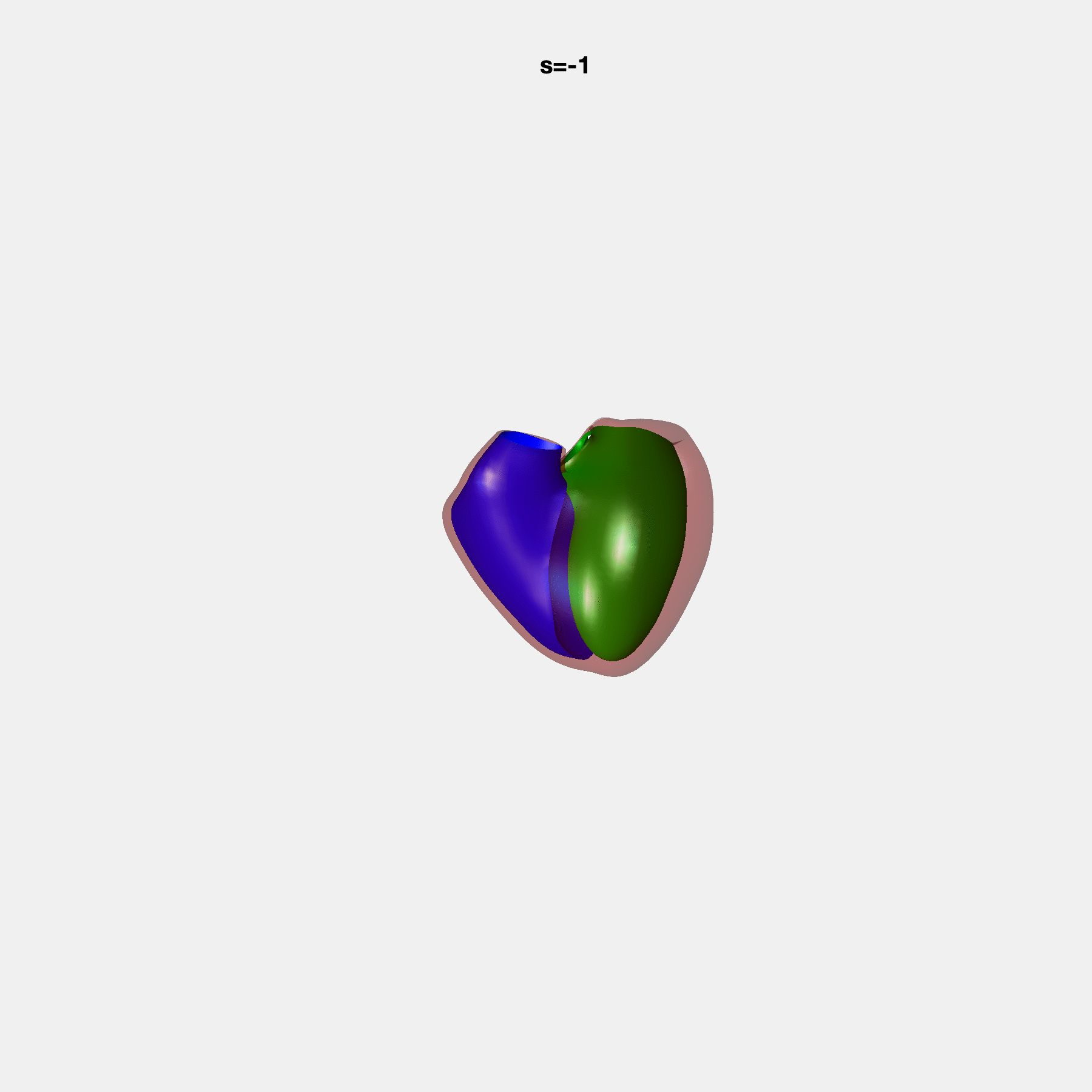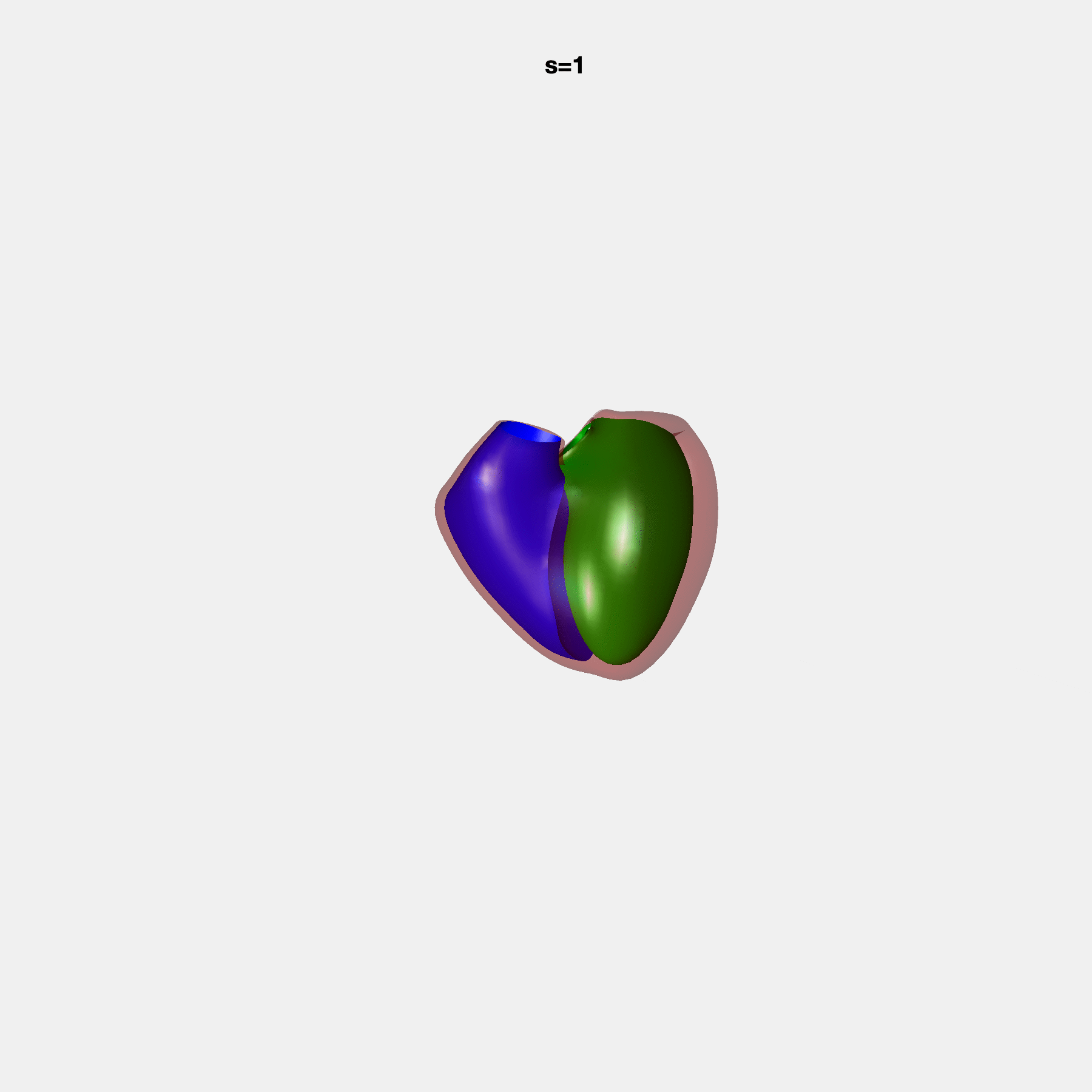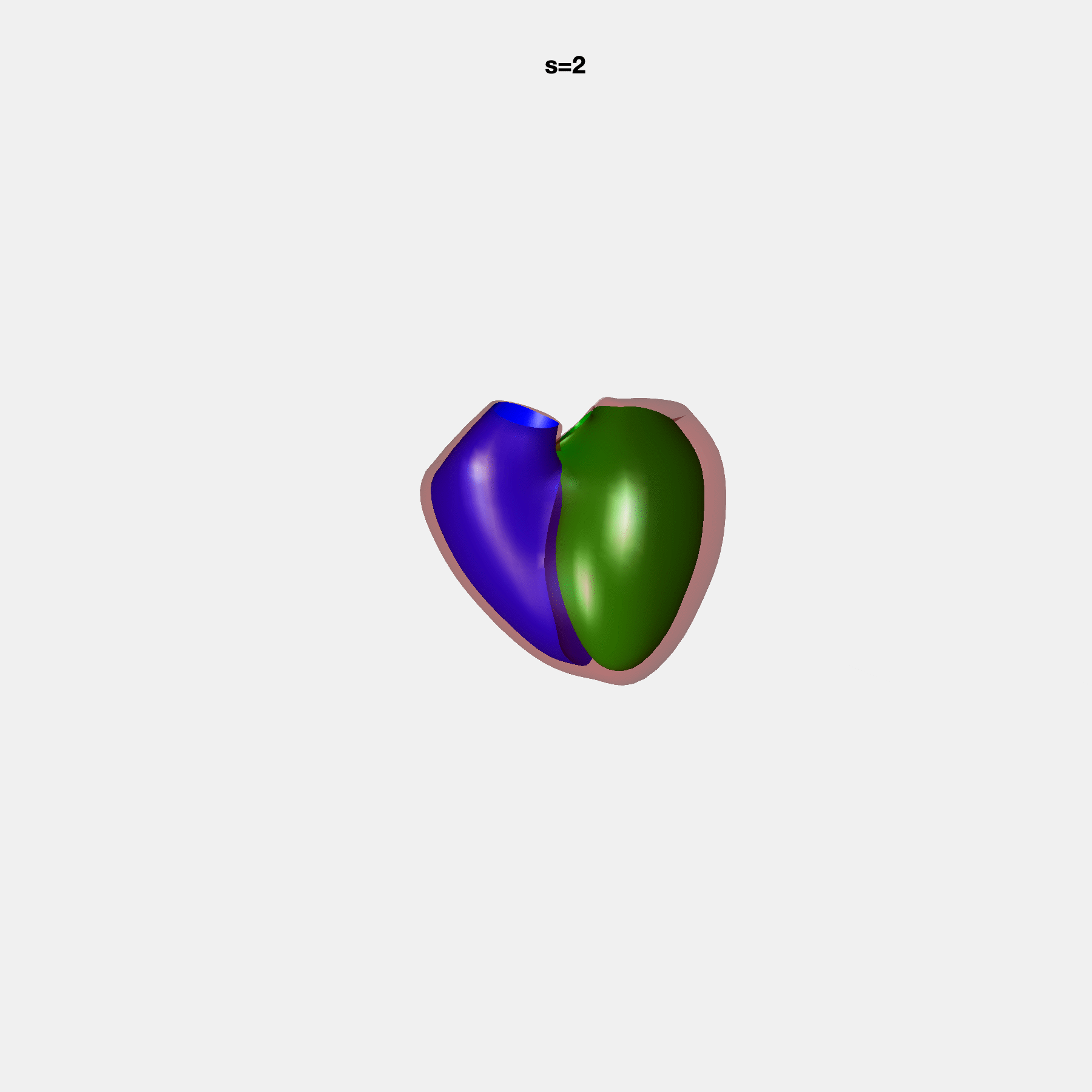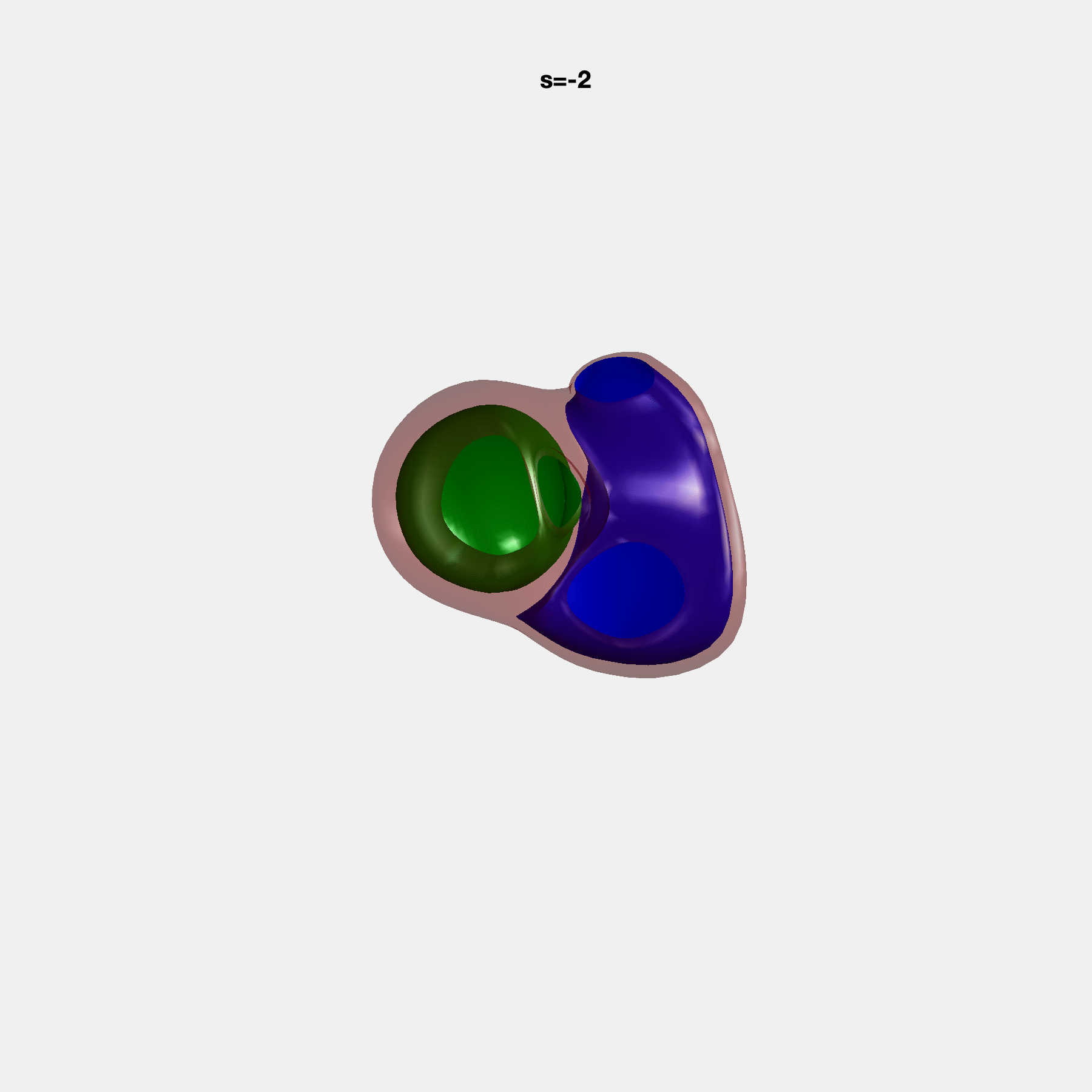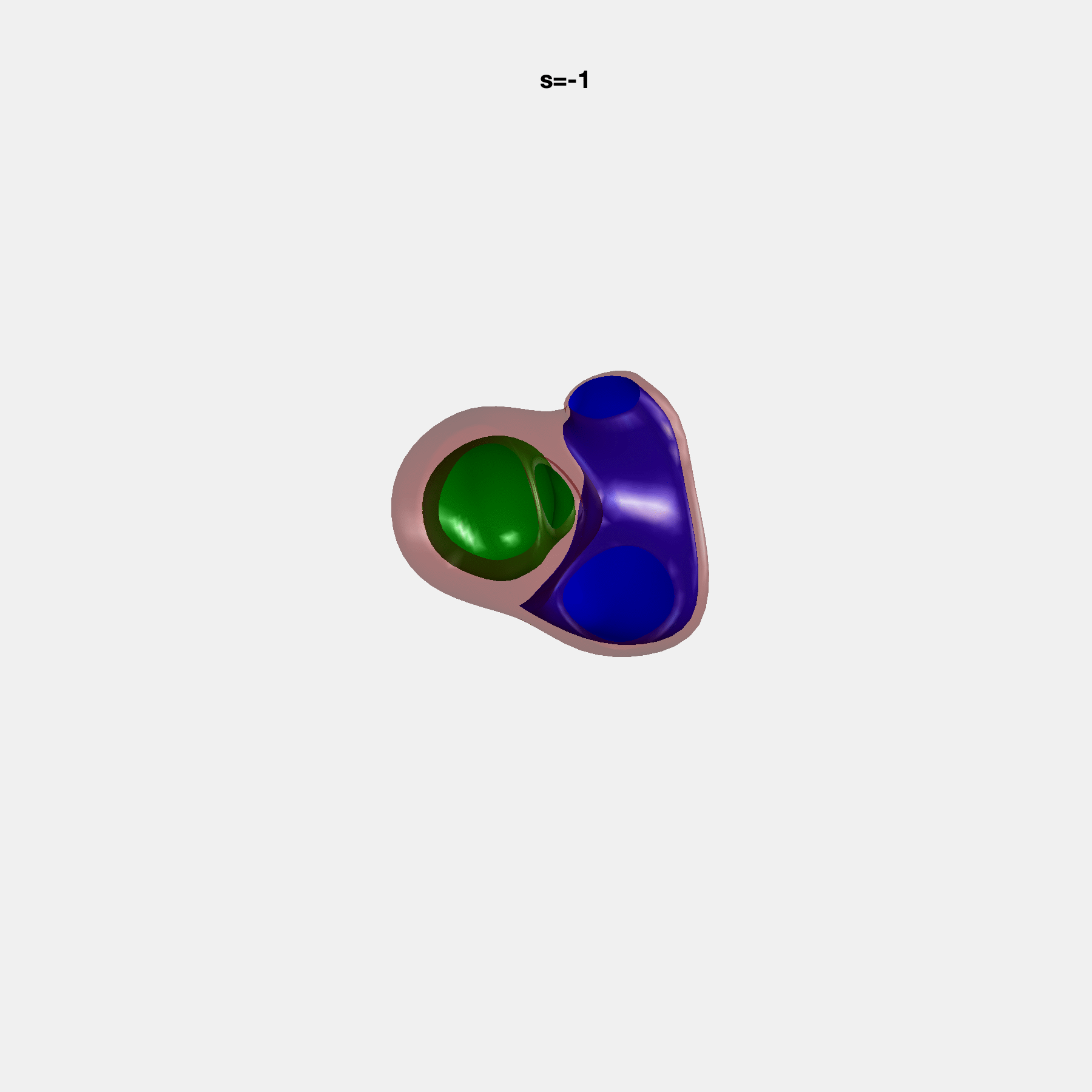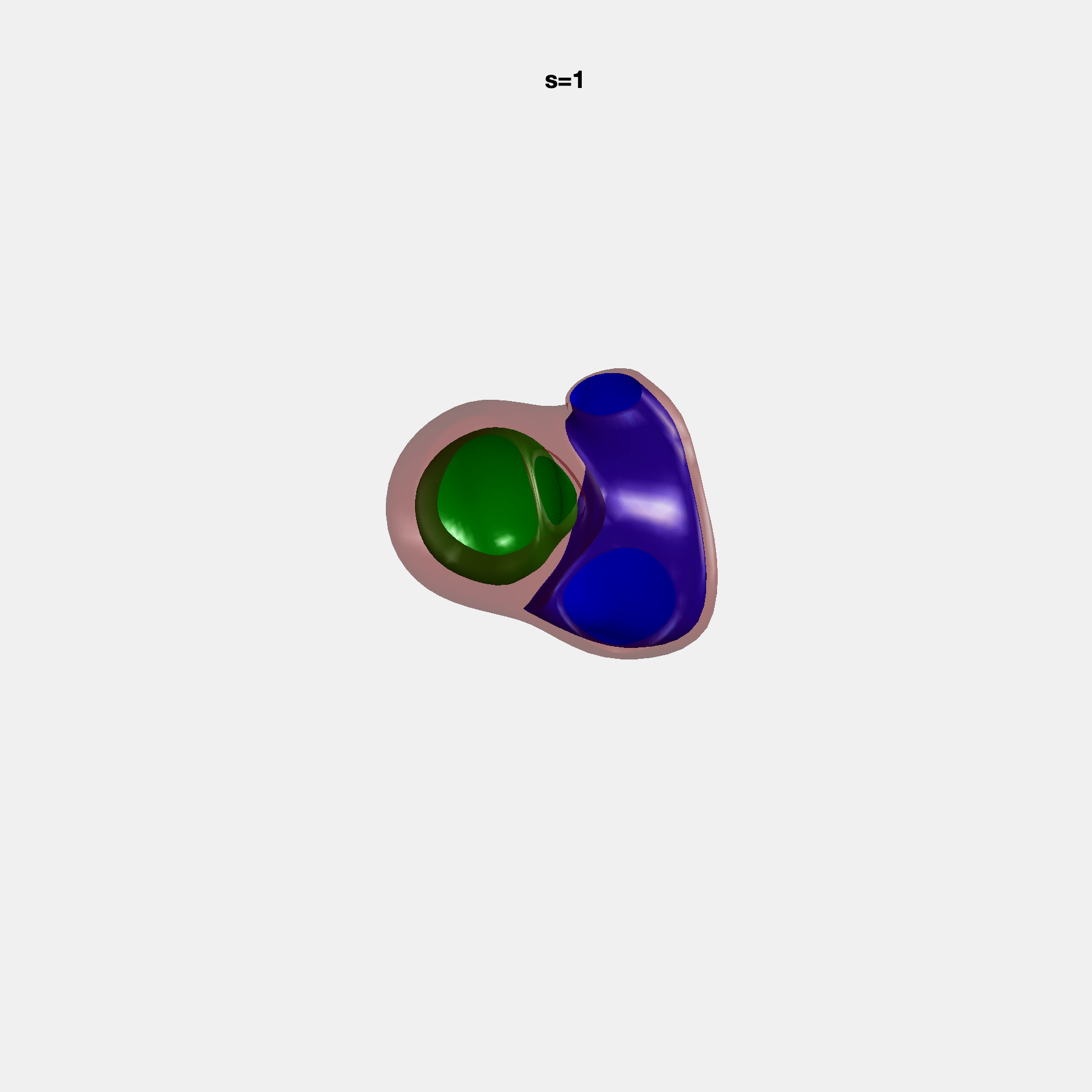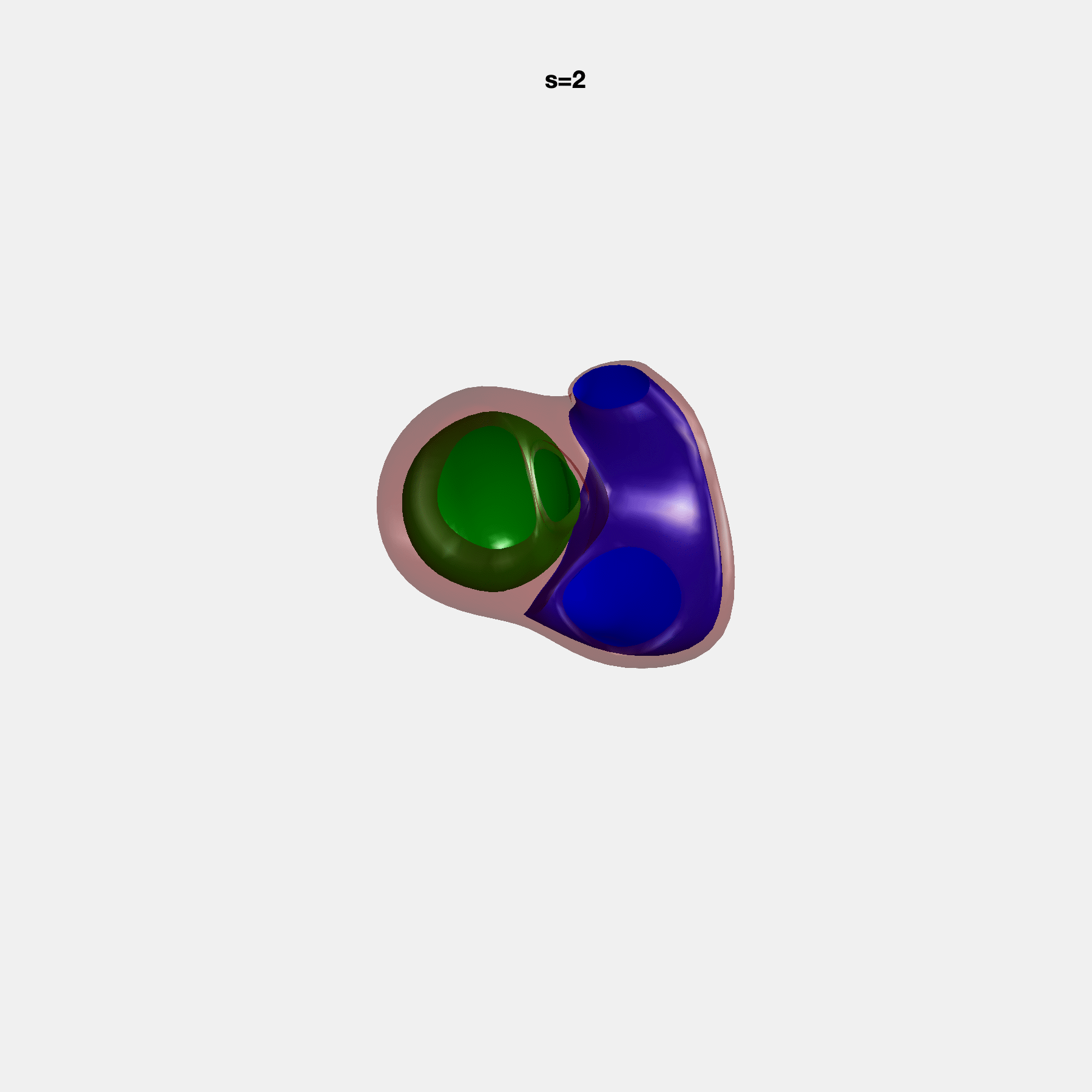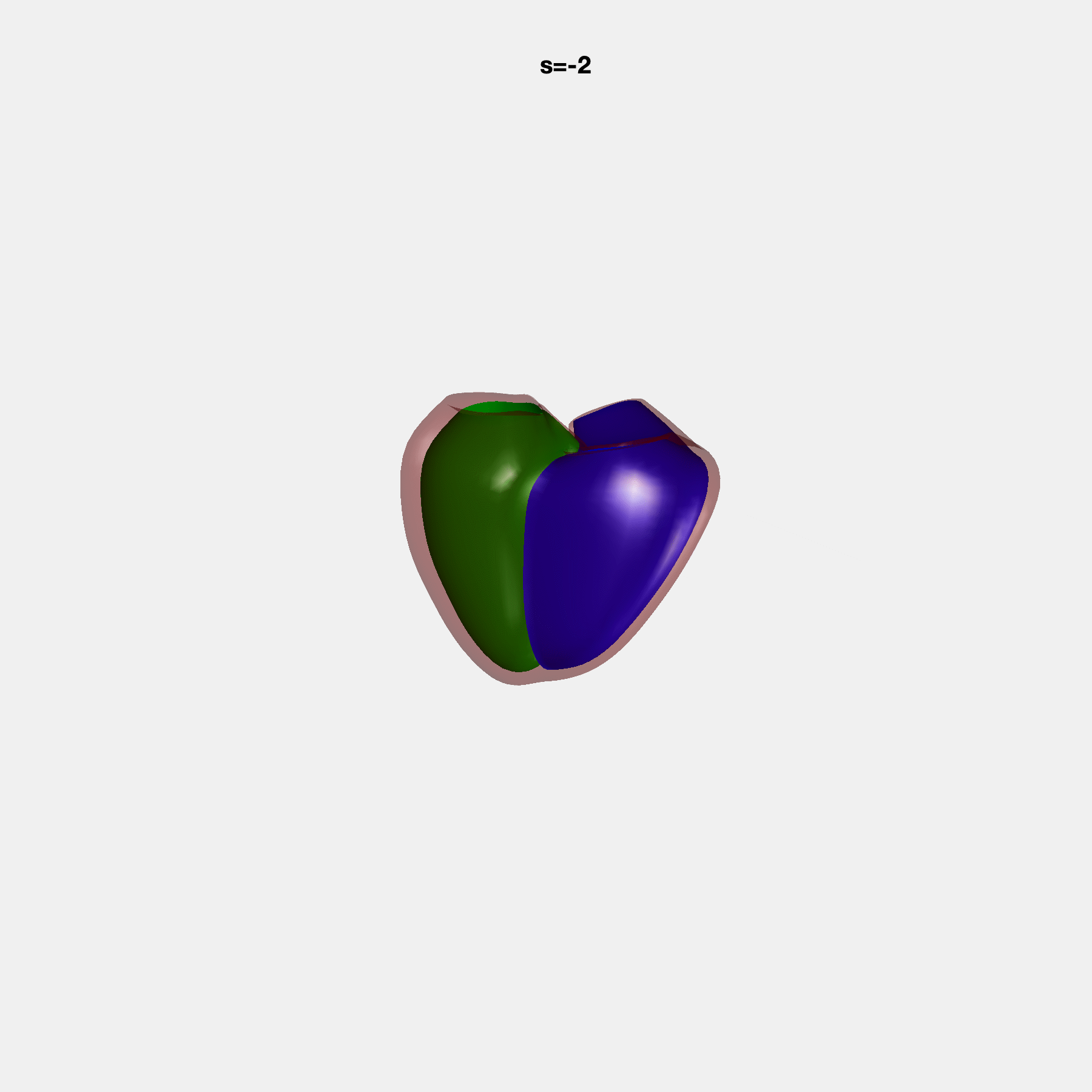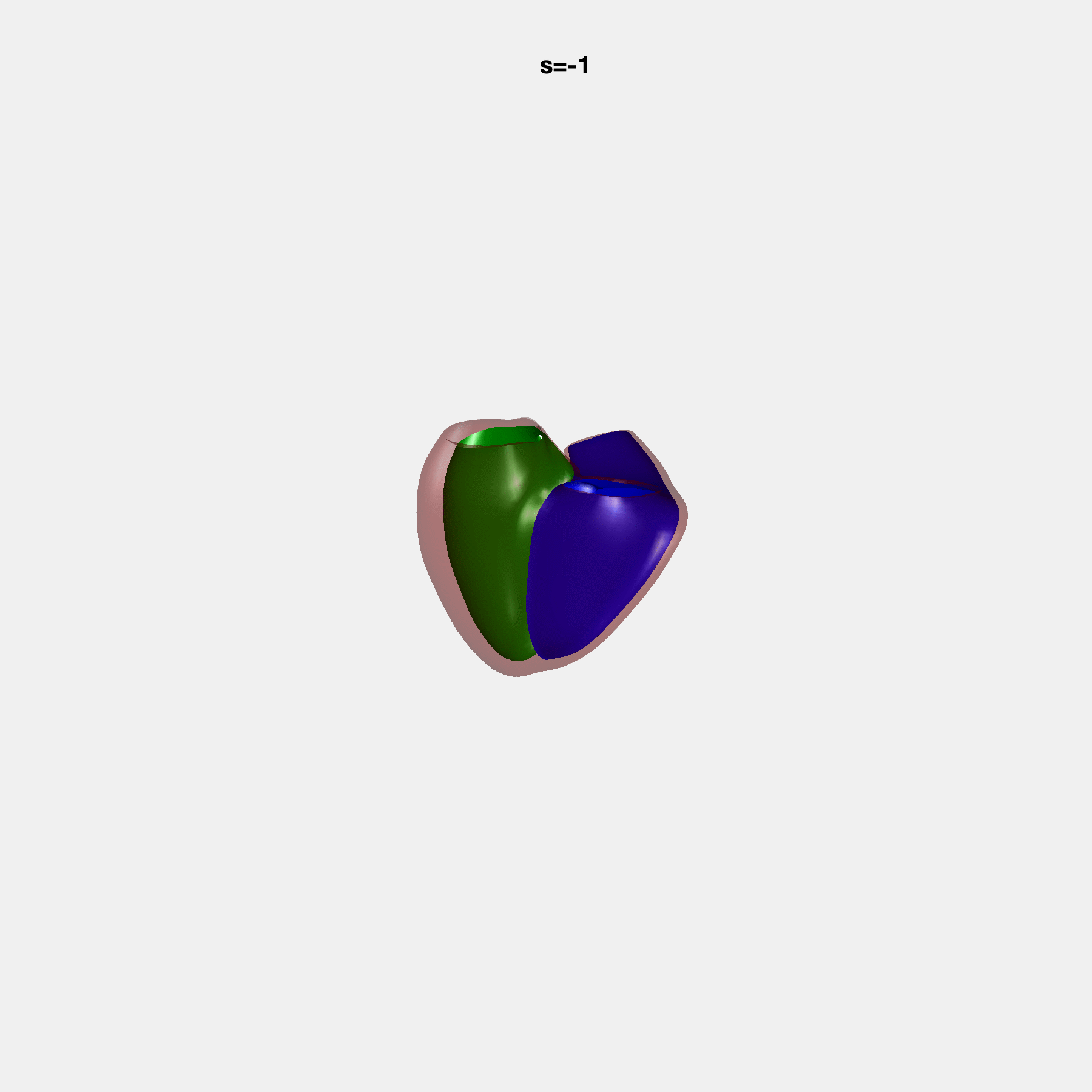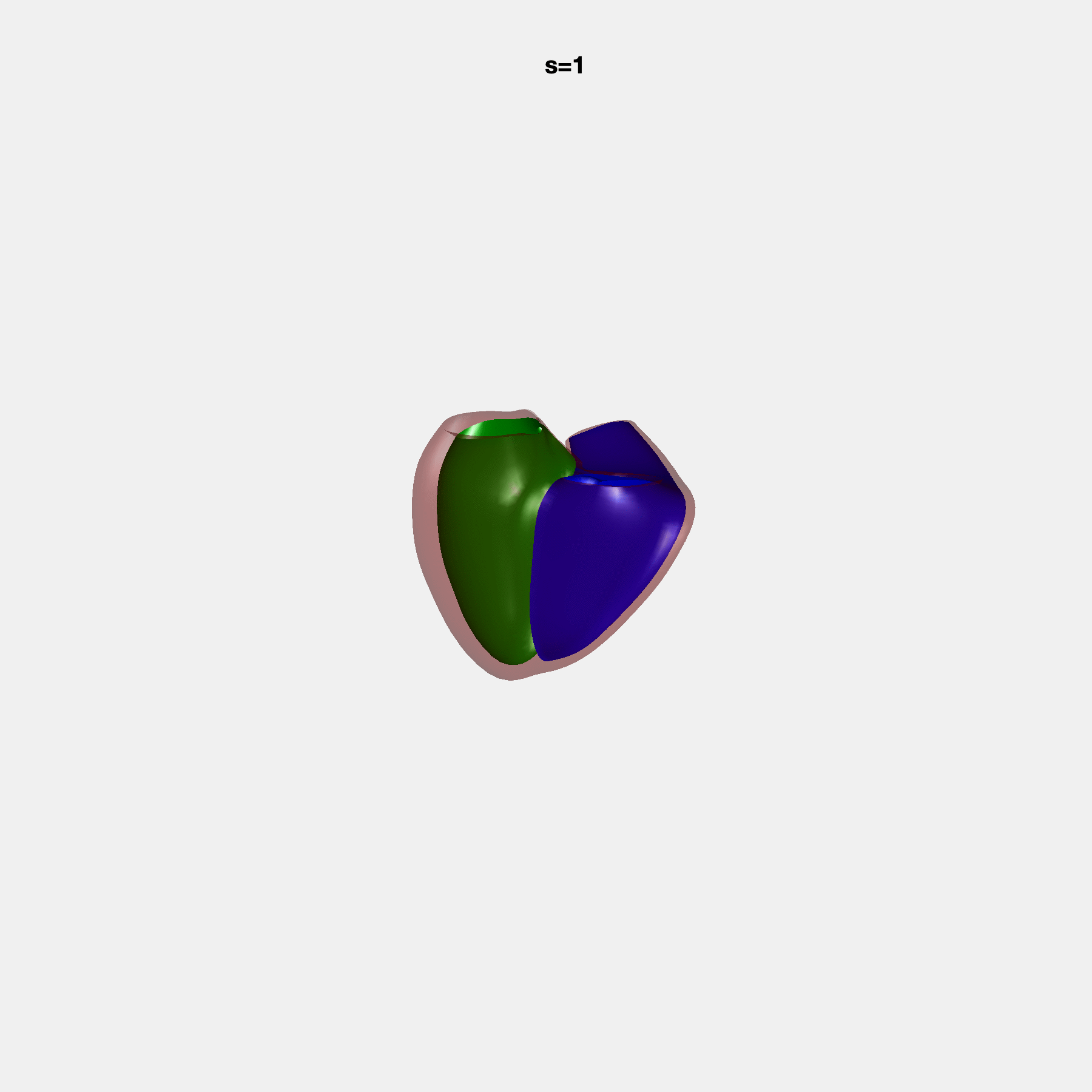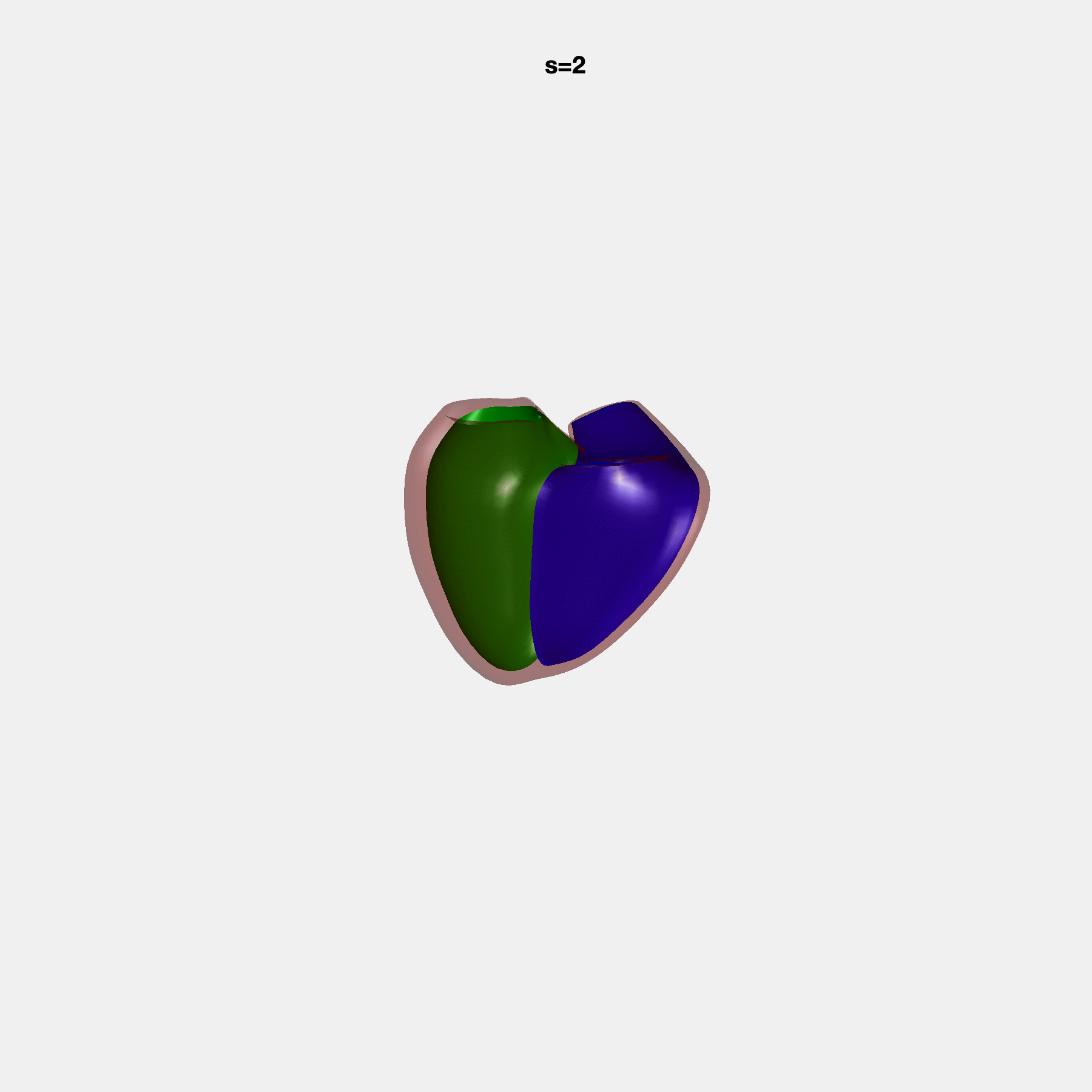Heart shape captures variation in cardiac structure beyond traditional phenotypes of mass and volume. Although observational studies have demonstrated associations with cardiometabolic risk factors and diseases, its genetic basis has not been investigated. We utilised cardiovascular magnetic resonance images from 38,858 UK Biobank participants to construct a heart shape atlas from bi-ventricular end-diastolic and end-systolic surface mesh models through principal component (PC) analysis.
Genome-wide association studies were performed on the first 14 PCs that captured 83.3% of shape variance. We identified the first 14 PCs with estimated biological features below:
| PC | % Shape variance captured | Estimated biological feature |
|---|---|---|
| 1 | 34.1 | Overall heart size |
| 2 | 10.6 | Mitral and tricuspid annular plane systolic excursion |
| 3 | 7.8 | Ventricular sphericalisation |
| 4 | 5.5 | LV basal position |
| 5 | 5.5 | Lateral-septal width |
| 6 | 4.2 | LV length |
| 7 | 4.0 | LV apical position |
| 8 | 2.4 | Tricuspid valve opening size |
| 9 | 2.1 | Systolic basal excursion |
| 10 | 2.0 | RV sphericity |
| 11 | 1.6 | Rounder ventricular apex |
| 12 | 1.3 | Relative LV size |
| 13 | 1.1 | Ejection fraction |
| 14 | 1.0 | LV size and reduced ejection fraction |
PCA Mode 2 showing variations in mitral and tricuspid annular plane systolic excursion.
PCA Mode 3 showing variations in ventricular sphericalisation.
PCA Mode 8 showing variations in the tricuspid valve opening size.
PCA Mode 9 showing variations in the systolic basal excursion.
PCA Mode 11 showing variations in the rounder ventricular apex.